A subscription to JoVE is required to view this content. Sign in or start your free trial.
Synthetic Condensates and Cell-Like Architectures from Amphiphilic DNA Nanostructures
In This Article
Summary
We present a protocol for preparing synthetic biomolecular condensates consisting of amphiphilic DNA nanostars starting from their constituent DNA oligonucleotides. Condensates are produced from either a single nanostar component or two components and are modified to sustain in vitro transcription of RNA from an embedded DNA template.
Abstract
Synthetic droplets and condensates are becoming increasingly common constituents of advanced biomimetic systems and synthetic cells, where they can be used to establish compartmentalization and sustain life-like responses. Synthetic DNA nanostructures have demonstrated significant potential as condensate-forming building blocks owing to their programmable shape, chemical functionalization, and self-assembly behavior. We have recently demonstrated that amphiphilic DNA "nanostars", obtained by labeling DNA junctions with hydrophobic moieties, constitute a particularly robust and versatile solution. The resulting amphiphilic DNA condensates can be programmed to display complex, multi-compartment internal architectures, structurally respond to various external stimuli, synthesize macromolecules, capture and release payloads, undergo morphological transformations, and interact with live cells. Here, we demonstrate protocols for preparing amphiphilic DNA condensates starting from constituent DNA oligonucleotides. We will address (i) single-component systems forming uniform condensates, (ii) two-component systems forming core-shell condensates, and (iii) systems in which the condensates are modified to support in vitro transcription of RNA nanostructures.
Introduction
Synthetic cells are micrometer-scale (10-50 µm) devices constructed from the bottom-up to replicate functions and structures of extant biological cells1,2. Synthetic cells are often bound by membranes constructed from lipid bilayer vesicles3,4,5,6,7, polymersomes8,9, or proteinosomes10,11, which can also be used to establish internal compartmentalisation12,13. Inspired by the membrane-less organelles known to sustain various functionalities in living cells14, structures such as polymer coacervates, biomolecular condensates, and hydrogels are gaining traction as versatile and robust alternatives to establish both external and internal compartmentalization in synthetic cells15,16,17,18.
Leveraging the versatile toolkit of DNA nanotechnology19, multiple solutions have been developed to engineer synthetic droplets and condensates from the self-assembly of artificial DNA nanostructures, whose size, shape, functionality, valency, and mutual interactions can be precisely programmed20. DNA droplets or condensates are biocompatible and can act as scaffolds for both synthetic cells and organelles, hosting chemical and biomolecular reactions21, computing information22,23, capturing and releasing cargoes24,25, and sustaining structural responses26.
Among the diverse designs of condensate-forming DNA nanostructures, amphiphilic DNA nanostars - dubbed C-stars - have proven robust and versatile27. C-stars are simple branched motifs consisting of a fixed DNA junction (typically four-way), from which double-stranded (ds)DNA arms emerge28. The arms are then tipped with hydrophobic moieties, typically cholesterol, rendering the nanostructures amphiphilic and driving their condensation following a straightforward one-pot annealing. C-star condensates afford precise structural and functional programmability, including the possibility of establishing multi-compartment architectures29,30, structurally responding to DNA and cation triggers31, synthesizing macromolecules29, capturing and releasing payloads32, and interacting with live cells33. Below, we will describe and discuss protocols to produce C-star condensates starting from their constituent oligonucleotides.
The protocol summarizes the preparation of unary (one-component) and binary (two-component) condensates, utilizing three different C-star designs (Figure 1) -"Non-responsive", "TMSD-responsive", and "RNA-templating". The "Non-responsive" C-star (panel A) consists of four "core strands" with distinct sequences forming the four-way junction. Four identical cholesterol-modified oligonucleotides are connected to the junction, ensuring that a cholesterol molecule is present at the end of each arm. The non-responsive C-stars constitute simple, inert scaffolds for unary and binary condensates. In the "TMSD-responsive" C-star (panel B), the connection between the cholesterolised strands and the junction is ensured by a "Toeholding bridge" strand, which features a dangling single-stranded (ss)DNA "toehold" domain. In the presence of an invader DNA strand with a complementary toehold domain, a toehold-mediated strand displacement reaction can be triggered34, whereby the invader displaces the Toeholding bridge, breaking the connection between the junction and the hydrophobic moieties and triggering the disassembly of the DNA network32. Finally, the "RNA templating" C-star (panel C) includes a "Base" modification complementary to a "Bridge" strand, the latter of which links the transcribable ssDNA template for the Broccoli aptamer29. Sequence details of the constituent oligonucleotides for the three types of C-star designs mentioned here can be found in Supplementary Table 1 and across previous works29,30,32.
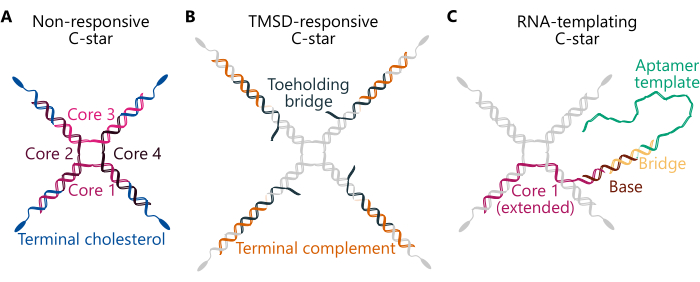
Figure 1. Schematics of three different designs of amphiphilic DNA nanostars (C-stars). Oligonucleotide sequences for various examples of the C-stars described here can be found in Supplementary Table 1. (A) Schematic of a C-star designed to form non-responsive condensates, with the component oligonucleotide strands "Core 1", "Core 2", "Core 3", "Core 4", (coloured in shades of pink) and "Terminal cholesterol" (coloured in blue). Each unique colour represents an oligonucleotide strand of unique sequence. "Core 1" and "Core 3" are each partially complementary to "Core 2" and "Core 4", but non-complementary to each other. (B) Schematic of a C-star designed to disassemble upon the addition of an invading strand via toehold-mediated strand displacement, as described in previous work32. This C-star is composed of "Core" and "Terminal cholesterol" strands (coloured in grey) as well as a "Terminal complement" (shown in orange) and a "Toeholding bridge" strand (shown in dark teal). The latter contains a six-nucleotide overhang to which an appropriately designed invader strand can bind and subsequently entirely displace the "Toeholding bridge" strand, which causes the dissociation of the central nanostar junction (composed of "Core 1, 2, 3, and 4") from the duplexes composed of the "Terminal complement" and "Terminal cholesterol" strands. (C) Schematic of a C-star functionalised with a DNA template for an RNA aptamer. This, too, is composed of the "Terminal cholesterol" strand and "Core 2, 3, and 4" (all shown in grey), as well as an extended version of the "Core 1" strand (shown in pink), a "Base" strand (brown), a "Bridge" strand (yellow), and the "Aptamer template" (green). The DNA duplex composed of the latter two strands forms the T7 polymerase promoter region, which marks the transcription start site. Please click here to view a larger version of this figure.
C-star condensates form upon thermal annealing of the constituent oligonucleotides, which in the protocol presented here is conducted within sealed glass capillaries with a high aspect ratio rectangular cross-section. These containers offer multiple key advantages: i) Sealing ensures that evaporation is completely prevented over the (sometimes slow) annealing steps; ii) The optical-quality flat bottom of the capillaries enables imaging of the self-assembly (or disassembly) transient; iii) the high aspect ratio of the capillaries ensures that heavy condensates settle over a wide, flat area, reducing chances of coalescence and aggregation at later stages of the self-assembly transient that would occur in wedge-shaped containers (e.g., microcentrifuge tubes), and producing relatively monodisperse condensate populations; iv) performing the annealing in an elongated glass capillary minimizes exposure of the sample to hydrophobic interfaces (air, plastic or oil), which have been observed to perturb self-assembly by recruiting the amphiphilic cholesterolised oligonucleotides. Once the assembly protocol is completed, condensates can be extracted from glass capillaries for further experiments that involve additional reagents.
Protocol
NOTE: The protocol is divided into three sections. Section 1 describes the prerequisite steps, including the preparation of DNA oligonucleotides and glass capillaries. Section 2 describes the preparation of C-star condensates of various designs, including one- and two-component designs, and their extraction from the glass capillaries. Section 3 describes the use of one-component RNA templating C-star condensates for the synthesis of an RNA aptamer. The user must follow good lab practice throughout, ensure that all necessary risk assessments and mitigations are in place, and wear appropriate personal protective equipment (PPE), including gloves, safety glasses, and a lab coat. The cleaning of glass capillary tubes requires their sonication, first in a surfactant solution and second in isopropanol or ethanol. The extraction of C-star condensates from capillary tubes requires the use of a diamond scribing pen to score and snap the glass, with an associated risk of injury from glass fragments. Key materials, equipment, and reagents used are listed in the Table of Materials. Most non-functionalized oligonucleotides are purified by the supplier using standard desalting, with the exception of the "extended Core 1" and "Aptamer template" strands, which are ordered with polyacrylamide gel electrophoresis (PAGE) purification. Cholesterol-modified oligonucleotides are purified by the supplier using reverse-phase high-performance liquid chromatography (HPLC).
1. Prerequisites
NOTE: The following solutions should be prepared in ultrapure (Type I) water and filtered using 0.22 µm syringe filters: Tris-EDTA (TE) buffer, comprising 10 mM Tris, 1 mM EDTA, at pH ~8.0; TE buffer supplemented with 2 M NaCl; and TE buffer supplemented with 0.3 M NaCl. Buffer solutions should be used within 2 weeks of preparation and stored at 4 °C when not in use. In addition, a 1 vol% solution of alkaline optical detergent in ultrapure water will be used for cleaning the glass capillaries.
- Preparation of DNA oligonucleotides from lyophilized state
- Briefly centrifuge (2000 x g for 10-30 s) the tubes of lyophilized DNA oligonucleotides to ensure pellets of lyophilized DNA collect at the bottom.
NOTE: The centrifugation is done at room temperature (RT) for 10-30 s. The mini centrifuge used here defaults to 2000 x g (6000 rpm). - Add the appropriate amount of TE buffer to reconstitute each oligonucleotide at 100 µM. Vortex thoroughly to ensure complete dissolution, then spin down tubes to collect all liquid as per step 1.1.1.
- For each stock solution, measure absorbance at 260 nm and calculate the final concentration using the extinction coefficient of the oligonucleotide sequence under test.
NOTE: The extinction coefficient for a given oligonucleotide sequence is usually found on the supplier's Data Sheet but can also be calculated with online tools using the nearest neighbor model35 or tabulated values of the extinction coefficient of individual nucleotides36. - Store the rehydrated oligonucleotides at 4 °C for short periods of time (up to 1 week) or at -20 °C for longer-term storage (up to 6 months).
- Briefly centrifuge (2000 x g for 10-30 s) the tubes of lyophilized DNA oligonucleotides to ensure pellets of lyophilized DNA collect at the bottom.
- Cleaning of glass capillary tubes
- Sonicate in 1% alkaline optical detergent for optical components in deionized water.
- Take the required number of glass capillary tubes to be cleaned and place them into a tall, narrow container (e.g., a beaker or a 15/50 mL centrifuge tube) such that the capillaries do not lie flat on the base of the container.
- Add the detergent to the container, filling it just above the level of the capillary tubes. Ensure there are no trapped air bubbles inside the capillary tubes - tap to remove, if necessary. Loosely cover the container (e.g., in tin foil or using the centrifuge tube lid, ensuring it is not tightly sealed).
- Set a sonicating water bath to 40 °C, place the covered container upright into the bath, and sonicate for 30 min. Ensure the foil, if used, does not come into contact with the water in the sonicating bath.
NOTE: The sonicating bath used in this study defaults to a sonication frequency of 60 Hz and a sonication power of 150 W.
CAUTION: Never add or remove items from a sonicating bath while running; always pause the protocol first. - Once sonication is complete, turn off the heating element of the sonicating water bath, then thoroughly rinse the capillaries in the container with deionized or ultrapure water - a minimum of five times, discarding the rinsing water each time.
- Sonicate in 1% alkaline optical detergent for optical components in deionized water.
- Sonicate in isopropanol or ethanol.
- To the glass capillary tubes in the cleaning container, add isopropanol or ethanol (minimum 70%), filling to just above the level of the capillary tubes. As in step 1.2.1.2, ensure there are no trapped air bubbles in the capillary tubes and loosely cover the container.
- Place the container upright into the sonicating bath and sonicate for 15-30 min.
CAUTION: Isopropanol and ethanol are flammable, with flash points below RT. Minimize the volumes of solvents used during cleaning by choosing a container of appropriate size, ensure that the containers are loosely covered, and do not leave the sonicating bath unattended during this stage. - Once sonication is complete, appropriately dispose of the isopropanol or ethanol and dry the capillaries under nitrogen, handling them with a lint-free tissue.
2. Preparation and extraction of C-star condensates (Figure 2)
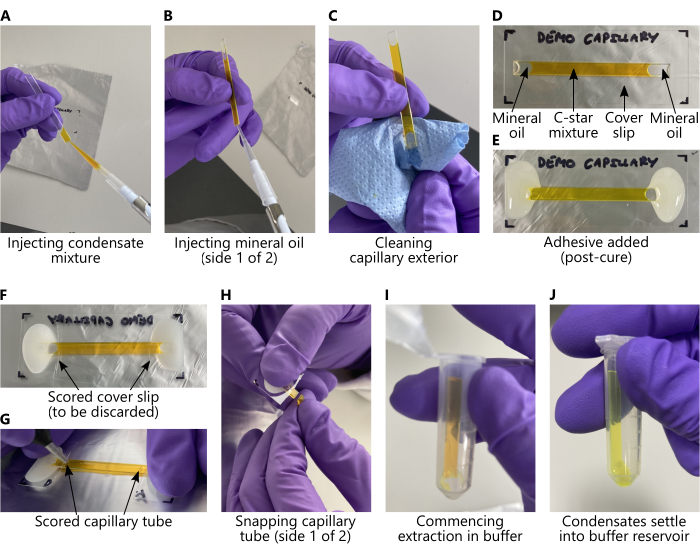

Figure 2: Loading C-star mixtures and extracting condensates from glass capillary tubes. In all panels, the C-star mixture has been replaced with an aqueous solution of 25 mM calcein to aid visibility. (A-E) Key steps, in order, to be taken prior to annealing, corresponding to protocol sections 2.1 and 2.2. (F-J) Key steps, in order, to be taken after annealing, corresponding to protocol section 2.3. During extraction (panels (I-J)), DNA condensates will sediment from the capillary into the buffer reservoir as long as the microcentrifuge tube is stored vertically. Condensates will not be visible to the naked eye. Please click here to view a larger version of this figure.
- Preparation of C-star mixtures for single-component condensates
NOTE: Sequence details for various designs of single components can be found in Supplementary Table 1 and across previous works29,30,32. The slow annealing protocols defined in steps 2.1.9 and 2.2.2 were developed to ensure the relaxation of the nanostar networks into compact morphologies, given the strongly temperature-dependent rheological properties of nanostar networks37.- Oligonucleotide and buffer components will vary depending on the C-star design desired. Prepare a 60 µL mixture of single-component C-star condensates at 5 µM concentration for either the non-responsive four-arm C-star, the TMSD-responsive C-star, or the RNA-templating C-star.
- For each design, pipette the appropriate components listed in Supplementary Table 2 into a separate microcentrifuge tube and mix thoroughly via pipetting.
NOTE: All oligonucleotide stock solutions are assumed to be at 100 µM concentration in TE. - Withdraw the entire 60 µL of C-star solution and pipette carefully to inject the mixture into the cleaned, dry glass capillary tube as shown in Figure 2A, taking care to avoid the introduction of air bubbles.
- Using a pipette, inject approximately 9-12 µL of mineral oil into each end of the capillary tube, capping the sample such that there is no free interface between the C-star solution and the air (see Figure 2B). Then, carefully dry the capillary tube with tissue paper to ensure no oil is present on the outside of it (Figure 2C), taking care not to wick mineral oil or C-star solution out of the capillary tube. The end result is demonstrated in Figure 2D.
- Prepare a small batch of two-part epoxy glue to adhere each end of the capillary tube, flat side down, to a glass cover slip. Ensure the openings of the tube are completely covered in glue, forming a continuous seal, as shown in Figure 2E.
- Set aside to cure for a minimum of 3 h, but preferably overnight.
- After ~30 min of curing, inspect the glue layer for the presence of gaps in the seal caused by air bubbles. If these are present, seal them using a small quantity of glue.
- Wrap the capillary tube glued to the coverslip in tin foil, ensuring the foil is kept flat at the underside of the glass cover slip. The foil ensures good thermal contact between the same and the heating element of the thermal cycler (see next step).
- Place the wrapped sample into a thermal cycler and anneal using the following protocol: Hold at 95 °C for 30 min, then cool from 85 °C to 50 °C at -0.04 °C·min-1, and then cool from 50 °C to RT at -0.5 °C·min-1.
- Store the annealed C-star condensates at 4 °C or RT for extended periods (months) as long as the capillary remains sealed, as the thermal annealing effectively sterilizes the sample and denatures nucleases. Keep the capillary flat during storage and handling to prevent condensates from sedimenting towards one end and aggregating.
- Preparation of C-star mixtures for binary condensates
NOTE: To inform the selection of C-star populations for binary systems, refer to the work by Malouf et al., which describes the design rules and expected phase behavior of binary C-star condensates30.- To prepare binary condensates, combine 30 µL volumes of the C-star mixtures described in step 2.1 above to a total volume of 60 µL, mix thoroughly, load into capillaries, and follow steps 2.1.4-2.1.8.
- Place the wrapped sample into a thermal cycler and anneal using the following protocol: Hold at 95 °C for 30 min, then cool from 85 °C to 40 °C at -0.01 °C·min-1, and then cool from 40 °C to RT at -0.1 °C·min-1.
- Extraction of C-star condensates from capillary tubes.
- Prepare a large (1.5 mL or 2.0 mL) microcentrifuge tube containing 60 µL of 0.3 M NaCl in TE buffer.
- Unwrap the capillary sample and use a diamond scribing pen to score the underside of the coverslip at each inner end of the glued area (Figure 2F). Break off the coverslip in this region and discard appropriately.
- Thoroughly clean the capillary tubes with ethanol and dry.
- Score each end of the capillary tubes with the diamond scribing pen, first with the underside (Figure 2G), then flipping and scoring the tubes "right side up".
- Ensure that the score lines do not overlap the oil-water interface. They must be within the aqueous region to prevent oil from remaining in the extracted sample.
- Snap off the ends of the capillary tubes (Figure 2H), retaining the central part of the tube and discarding the rest. Place the cut capillary tube into the microcentrifuge tube prepared in step 2.3.1, ensuring that the bottom of the capillary tube makes contact with the buffer solution (Figure 2I).
- The condensates will sediment from the tube into the buffer reservoir through gravity; allow this to take place for a minimum of 10 min (Figure 2J). Remove the cut capillary tube and discard it appropriately. This gentle extraction approach limits the risk of condensate aggregation.
3. Transcription of an RNA aptamer from RNA-templating C-star condensates
NOTE: For the production of the Broccoli RNA aptamer, a solution of difluoro-4-hydroxybenzylidene imidazolidinone (DFHBI) is required - DFHBI powder is first prepared as a stock solution at 10 mM in dimethyl sulfoxide (DMSO), which is then diluted to 600 µM in RNase- and DNase- free water.
- Washing RNA-templating C-star condensates
NOTE: RNA-templating C-star condensates are washed three times to ensure unbound template oligonucleotides are removed.- Allow the solution of extracted condensates to settle for at least 5 min, and then remove approximately half the volume of the supernatant.
- For a 60 µL volume of extracted condensates, remove between 25-30 µL of the supernatant.
- Pipette from the top of the liquid level to minimize the number of condensates removed during this step.
- Add the same volume of 0.3 M NaCl in TE to replace the removed supernatant and mix by pipetting.
- Repeat the previous two steps for a total of three cycles.
- Allow the solution of extracted condensates to settle for at least 5 min, and then remove approximately half the volume of the supernatant.
- Preparation of T7 transcription mixture
NOTE: The T7 transcription mixture is prepared using the CellScript T7-FlashScribe Transcription Kit. Step 3.2.2 is a modification of the manufacturer's protocol, which can be found here38. Here we describe the transcription of the Broccoli RNA aptamer, which induces fluorescence in DFHBI upon binding. For other light-up aptamers, replace the volume of DFHBI described in step 3.2.2 with the appropriate fluorogen. For other transcripts, remove this component entirely.- Prepare a stock solution of 10 mM DFHBI in DMSO, then dilute an aliquot to an end concentration of 600 µM using RNase- and DNase- free water.
- Defrost the transcription kit components on ice and then pipette the following into an autoclaved microcentrifuge tube at room temperature using sterile pipette tips: 2 µL of 10x T7-Transcription Buffer (provided in the transcription kit), 1.8 µL of 100 mM ATP, 1.8 µL of 100 mM CTP, 1.8 µL of 100 mM UTP, 1.8 µL of 100 mM GTP, 2 µL of 100 mM DTT, 2 µL of 600 µM DFHBI, 0.5 µL of RNase inhibitor, 2 µL of T7-enzyme solution (provided in the transcription kit).
- Gently mix the solution by pipetting.
- Use transcription mixture for step 3.3 (synthesis of RNA transcript) immediately after preparing.
- Synthesis of RNA transcript
- Into a chamber suitable for microscopy imaging, pipette 3.3 µL of the washed condensates prepared from RNA-templating C-stars and add the total volume of the transcription mixture prepared prior.
- Acquire microscopy images for a duration of 18 h, starting immediately after adding the transcription mixture to the condensates.
NOTE: Microscopy settings will depend on the system prepared. Ensure that the appropriate excitation and emission settings are used for any fluorophores in the sample, along with exposure times that will not lead to saturation as transcription progresses. As for any time-lapse, a compromise should be found between a high enough laser or LED power for a good signal while also minimising the risk of photobleaching. Photobleaching over long periods of time will be minimised on LED-based systems compared to laser-based systems. Suggested imaging intervals: 20 min for the first 2 h, and 30 min subsequently.
Results
After annealing, C-star condensates can be imaged directly in the capillary tube, or after extraction, to confirm their formation. For all C-star design variations, one should observe distinct spherical or polyhedral condensates approximately 10-50 µm in diameter, the latter forming when crystallization occurs28,32. For single-component condensates, the condensates should be discrete and uniform in appearance, and may appear polyhedral (...
Discussion
The protocol described here provides an approach for the preparation of one- or two- component condensates from amphiphilic DNA nanostars, with design variations to introduce different responses into the condensates. The given protocol produces condensates in a buffer solution of 0.3 M NaCl in TE, but the buffer conditions can be amended by appropriately modifying the volumes listed above. Previous work has studied the formation of C-star condensates in 0.2 M NaCl in TE and 0.1 M NaCl in TE and in phosphate-buffered sali...
Disclosures
No conflicts of interest declared.
Acknowledgements
LM, LDM, and DT acknowledge support from the European Research Council (ERC) under the Horizon 2020 Research and Innovation Programme (ERC-STG No 851667 - NANOCELL). LDM acknowledges support from a Royal Society Research Grant for Research Fellows (RGF/R1/180043) and support from a Royal Society University Research Fellowship (UF160152, URF/R/221009).
Materials
| Name | Company | Catalog Number | Comments |
| 0.22 μm syringe filters | Sigma-Aldrich | SLGVR33RB | |
| 24 x 60 mm #1.5 Rectangular cover glasses, Menzel Gläser | VWR | 631-0853 | |
| 2-Propanol | Sigma-Aldrich | 34683 | |
| 6 L Ultrasonic Cleaner with Digital Timer and Heat, 230 VAC | Cole-Parmer | WZ-08895-11 | |
| Araldite Rapid Adhesive 2 Part Epoxy Glue | RS | ARA-400005 | |
| Bio-Rad C1000 thermal cycler | Bio-Rad | 1851197 | |
| Brand Microcentrifuge Tube 2 mL with Locking Lid | Fisher Scientific | 15338665 | 2 mL microcentrifuge tubes for the extraction of C-star condensates |
| Diamond Scribing Pen | RS | 394-217 | |
| Difluoro-4-hydroxybenzylidene imidazolidinone (DFHBI) | Sigma-Aldrich | SML1627 | |
| Dimethyl sulfoxide (DMSO) | Sigma-Aldrich | 472301 | |
| Eppendorf PCR Clean Colorless Safe-Lock Centrifuge Tubes | Fisher Scientific | 0030123301 | 0.5 mL microcentrifuge tubes for the preparation of C-star mixtures |
| Ethanol Absolute 99.8+% | Fisher Scientific | 10437341 | 70% ethanol is sufficient for cleaning purposes |
| Fisherbrand ZX4 IR Vortex Mixer | Fisherbrand | 13284769 | |
| Hellmanex III | Hellma | 9-307-011-4-507 | |
| Hollow Rectangle Capillaries ID 0.40 x 4.00 mm, 50 mm in length | CM Scientific | 2540-50 | |
| Mineral oil | Sigma-Aldrich | 69794 | |
| Mini Centrifuge, 230 V | PRISM(TM) | Z763128 | |
| NaCl | Sigma-Aldrich | S3014 | |
| NanoDrop One Spectrophotometer | Thermo Fisher Scientific | ND-ONE-W | Used to measure absorbance of oligonucleotides for concentration calculations |
| Oligonucleotides | Integrated DNA Technologies | Custom | Oligonucleotide sequences are unique to the C-star design required. |
| ScriptGuard RNase inhibitor | CELLSCRIPT | C-SRI6310K | RNase inhibitor |
| T7-FlashScribe Transcription Kit | Cambio | C-ASF3507 | |
| Tris-EDTA buffer, 100x stock solution | Sigma-Aldrich | 574793 | |
| UltraPure DNase/RNase-Free Distilled Water | Invitrogen | 10977035 | |
| VWR Spec-Wipe 3 Wipers | VWR | 21914-758 |
References
- Buddingh', B. C., Hest, J. C. M. v. Artificial cells: Synthetic compartments with life-like functionality and adaptivity. Acc Chem Res. 50 (4), 769-777 (2017).
- Fanalista, F., et al. Shape and size control of artificial cells for bottom-up biology. ACS Nano. 13 (5), 5439-5450 (2019).
- Dora Tang, T. -. Y., et al. Fatty acid membrane assembly on coacervate microdroplets as a step towards a hybrid protocell model. Nat Chem. 6 (6), 527-533 (2014).
- Deshpande, S., et al. Spatiotemporal control of coacervate formation within liposomes. Nat Commun. 10 (1), 1800 (2019).
- Rubio-Sánchez, R., et al. Thermally driven membrane phase transitions enable content reshuffling in primitive cells. J Am Chem Soc. 143 (40), 16589-16598 (2021).
- Jahnke, K., Huth, V., Mersdorf, U., Liu, N., Göpfrich, K. Bottom-up assembly of synthetic cells with a DNA cytoskeleton. ACS Nano. 16 (5), 7233-7241 (2022).
- Tran, M. P., et al. A DNA segregation module for synthetic cells. Small. 19 (13), 2202711 (2023).
- Mason, A. F., Buddingh', B. C., Williams, D. S., Hest, J. C. M. v. Hierarchical self-assembly of a copolymer-stabilized coacervate protocell. J Am Chem Soc. 139 (48), 17309-17312 (2017).
- Gumz, H., et al. Toward functional synthetic cells: In-depth study of nanoparticle and enzyme diffusion through a cross-linked polymersome membrane. Adv Sci. 6 (7), 1801299 (2019).
- Huang, X., Patil, A. J., Li, M., Mann, S. Design and construction of higher-order structure and function in proteinosome-based protocells. J Am Chem Soc. 136 (25), 9225-9234 (2014).
- Booth, R., Qiao, Y., Li, M., Mann, S. Spatial positioning and chemical coupling in coacervate-in-proteinosome protocells. Angew Chem Int Ed Engl. 58 (27), 9120-9124 (2019).
- Hindley, J. W., et al. Light-triggered enzymatic reactions in nested vesicle reactors. Nat Commun. 9 (1), 1093 (2018).
- Zubaite, G., Hindley, J. W., Ces, O., Elani, Y. Dynamic reconfiguration of subcompartment architectures in artificial cells. ACS Nano. 16 (6), 9389-9400 (2022).
- Hirose, T., et al. A guide to membraneless organelles and their various roles in gene regulation. Nat Rev Mol Cell Biol. 24 (4), 288-304 (2023).
- Guindani, C., Silva, L. C. d., Cao, S., Ivanov, T., Landfester, K. Synthetic cells: From Simple Bio-Inspired Modules to Sophisticated Integrated Systems. Angew Chem Int Ed Engl. 61 (16), e202110855 (2022).
- Adamala, K. P., et al. Present and future of synthetic cell development. Nat Rev Mol Cell Biol. 25 (3), 162-167 (2023).
- Allen, M. E., et al. Biomimetic behaviors in hydrogel artificial cells through embedded organelles. Proc Natl Acad Sci U S A. 120 (35), e2307772120 (2023).
- Cook, A. B., Novosedlik, S., Hest, J. C. M. v. Complex coacervate materials as artificial cells. Acc Mater Res. 4 (3), 287-298 (2023).
- Seeman, N. C., Sleiman, H. F. DNA nanotechnology. Nat Rev Mater. 3, 17068 (2018).
- Takinoue, M. DNA droplets for intelligent and dynamical artificial cells: from the viewpoint of computation and non-equilibrium systems. Interface Focus. 13 (5), 20230021 (2023).
- Liu, W., Lupfer, C., Samanta, A., Sarkar, A., Walther, A. Switchable hydrophobic pockets in DNA protocells enhance chemical conversion. J Am Chem Soc. 145 (13), 7090-7094 (2023).
- Wilner, O. I., Willner, I. Functionalized DNA nanostructures. Chem Rev. 112 (4), 2528-2556 (2012).
- Gong, J., Tsumura, N., Sato, Y., Takinoue, M. Computational DNA droplets recognizing miRNA sequence inputs based on liquid-liquid phase separation. Adv Funct Mater. 32, 2202322 (2022).
- Jeon, B. -. J., Nguyen, D. T., Saleh, O. A. Sequence-controlled adhesion and microemulsification in a two-phase system of DNA liquid droplets. J Phys Chem. 124 (40), 8888-8895 (2020).
- Sato, Y., Sakamoto, T., Takinoue, M. Sequence-based engineering of dynamic functions of micrometer-sized DNA droplets. Sci Adv. 6 (23), 3471 (2020).
- Saleh, O. A., et al. Vacuole dynamics and popping-based motility in liquid droplets of DNA. Nat Commun. 14 (1), 3574 (2023).
- Rubio-Sánchez, R., Fabrini, G., Cicuta, P., Michele, L. D. Amphiphilic DNA nanostructures for bottom-up synthetic biology. Chem Commun. 57 (95), 12725-12740 (2021).
- Brady, R. A., Brooks, N. J., Cicuta, P., Di Michele, L. Crystallization of amphiphilic DNA C-Stars. Nano Lett. 17 (5), 3276-3281 (2017).
- Leathers, A., et al. Reaction-diffusion patterning of DNA-based artificial cells. J Am Chem Soc. 144 (38), 17468-17476 (2022).
- Malouf, L., et al. Sculpting DNA-based synthetic cells through phase separation and phase-targeted activity. Chem. 9 (11), 3347-3364 (2023).
- Fabrini, G., Minard, A., Brady, R. A., Antonio, M. D., Michele, L. D. Cation-responsive and photocleavable hydrogels from noncanonical amphiphilic DNA nanostructures. Nano Lett. 22 (2), 602-611 (2022).
- Brady, R. A., Brooks, N. J., Foderà, V., Cicuta, P., Di Michele, L. Amphiphilic-DNA platform for the design of crystalline frameworks with programmable structure and functionality. J Am Chem Soc. 140 (45), 15384-15392 (2018).
- Walczak, M., et al. Responsive core-shell DNA particles trigger lipid-membrane disruption and bacteria entrapment. Nat Commun. 12 (1), 4743 (2021).
- Zhang, D. Y., Winfree, E. Control of DNA strand displacement kinetics using Toehold exchange. J Am Chem Soc. 131 (47), 17303-17314 (2009).
- . Integrated DNA Technologies OligoAnalyzer Tool Available from: https://www.idtdna.com/pages/products/custom-dna-rna/dna-oligos/custom-dna-oligos (2024)
- Cavaluzzi, M. J., Borer, P. N. Revised UV extinction coefficients for nucleoside-5'-monophosphates and unpaired DNA and RNA. Nucleic Acids Res. 32 (1), e13 (2004).
- Lattuada, E., Caprara, D., Piazza, R., Sciortino, F. Spatially uniform dynamics in equilibrium colloidal gels. Sci Adv. 7 (49), (2021).
- . T7-FlashScribeTM Transcription Kit Available from: https://www.cellscript.com/products/018pl0617CS.pdf (2017)
- Zadeh, J. N., et al. NUPACK: Analysis and design of nucleic acid systems. J Comput Chem. 32 (1), 170-173 (2011).
- Walczak, M., Brady, R. A., Leathers, A., Kotar, J., Di Michele, L. Influence of hydrophobic moieties on the crystallization of amphiphilic DNA nanostructures. The Journal of Chemical Physics. 158 (8), 084501 (2023).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved