A subscription to JoVE is required to view this content. Sign in or start your free trial.
Identification of Quiescent Cells in a Zebrafish T-Cell Acute Lymphoblastic Leukemia Model Using Cell Proliferation Staining
In This Article
Summary
We used cell proliferation staining to identify quiescent cells in the zebrafish T-acute lymphoblastic leukemia model. The stain is retained in non-dividing cells and reduced during cell proliferation, enabling the selection of dormant cells for further interrogation. This protocol provides a functional tool to study self-renewal in the context of cellular quiescence.
Abstract
Cellular quiescence is a state of growth arrest or slowed proliferation that is described in normal and cancer stem cells (CSCs). Quiescence may protect CSCs from antiproliferative chemotherapy drugs. In T-cell acute lymphoblastic leukemia (T-ALL) patient-derived xenograft (PDX) mouse models, quiescent cells are associated with treatment resistance and stemness. Cell proliferation dyes are popular tools for the tracking of cell division. The fluorescent dye is covalently anchored into amine groups on the membrane and macromolecules inside the cell. This allows for the tracking of labeled cells for up to 10 divisions, which can be resolved by flow cytometry.
Ultimately, cells with the highest proliferation rates will have low dye retention, as it will be diluted with each cell division, while dormant, slower-dividing cells will have the highest retention. The use of cell proliferation dyes to isolate dormant cells has been optimized and described in T-ALL mouse models. Complementary to the existing mouse models, the rag2:Myc-derived zebrafish T-ALL model provides an excellent venue to interrogate self-renewal in T-ALL due to the high frequency of leukemic stem cells (LSCs) and the convenience of zebrafish for large-scale transplant experiments.
Here, we describe the workflow for the staining of zebrafish T-ALL cells with a cell proliferation dye, optimizing the concentration of the dye for zebrafish cells, passaging successfully stained cells in vivo, and the collection of cells with varying levels of dye retention by live cell sorting from transplanted animals. Given the absence of well-established cell surface makers for LSCs in T-ALL, this approach provides a functional means to interrogate quiescent cells in vivo. For representative results, we describe the engraftment efficiency and the LSC frequency of high and low dye-retaining cells. This method can help investigate additional properties of quiescent cells, including drug response, transcriptional profiles, and morphology.
Introduction
Adult stem cells are responsible for the regeneration of differentiated cell types in a given organ and are predominantly present in a dormant, non-dividing state1,2. For example, hematopoietic stem cells (HSCs), which maintain the blood, largely remain quiescent, and only a small fraction enters the cell cycle to self-renew or differentiate to generate mature blood components3. Similarly, in cancers, a rare subpopulation of cells called cancer stem cells (CSCs) possess the self-renewal ability and are responsible for the long-term maintenance of the malignancy4. Cancer stem cells exist in vivo in a state of quiescence or slow growth, which may enable them to escape the anti-proliferative cancer treatments5, evade clearance by the immune system6, reduce oxidative stress, and enhance their DNA repair pathways7. Even a low number of CSCs left behind after treatment can potentially repopulate the tumor, resulting in a patient's relapse8. Accordingly, understanding cellular quiescence holds great promise for the identification of potential vulnerabilities of CSCs and the development of new ways to target them.
Cell proliferation dyes, such as the carboxyfluorescein succinimidyl ester (CFSE) stain and its derivatives, are commonly used to track the frequency of cell divisions9. The dye permeates across the cell membrane and, once inside the cell, undergoes activation by intracellular esterases into a fluorescent product. The resultant fluorescent compound is retained inside the cell through the covalent amide bonds formed between the succinimidyl moiety and the amine functional groups of intracellular proteins10. With each cellular division, the fluorescent compound is divided equally between the two resulting cells, causing a two-fold signal dilution. This dye enables the detection of up to 10 cell divisions through flow cytometry analysis11.
This approach has been previously utilized to enrich CSC populations in vitro by identifying slow-cycling populations of cells with high retention of the dye11,12. In T-ALL, CFSE has been used to track tumor growth in vivo in patient-derived xenografts in mice. Following cell labeling and three weeks of transplant, flow cytometry analysis showed a rare population of cells that still retained CFSE fluorescence. This population was associated with stemness, treatment resistance, and high similarity to relapse-causing cells in patients13. Accordingly, this dye provides a useful tool for the study of leukemia stem cell (LSC) phenotypes in T-ALL.
The aim of the work is to extend the application of the cell proliferation dye to study quiescence in vivo using a zebrafish T-ALL model. In particular, the rag2:Myc-driven zebrafish T-ALL model14 provides an excellent venue for the study of self-renewal due to the high frequency of LSCs compared to mouse models and human disease15. In addition, the use of zebrafish allows for large-scale transplantation studies, which can be done at a much lower cost of care and maintenance compared to their mouse counterparts16. Zebrafish are also excellent for live imaging applications, as fluorescently labeled tumor cells can be readily viewed using a simple fluorescence microscope to estimate the rate of tumor development16.
In this protocol, we describe the workflow for staining zebrafish T-ALL cells with cell proliferation dye followed by in vivo propagation of stained cells in syngeneic CG1 zebrafish. Upon leukemia development, we describe the sorting of cells that retained the dye and their use for a subsequent limiting dilution transplantation experiment to quantify rates of LSC self-renewal. This protocol can be extended for additional applications, including in vivo drug screening of potential compounds for the targeting of quiescent LSCs. In addition, collected cells can be used for different downstream analyses, such as transcriptomic profiling, proteomics, and metabolomics, offering unique insights into the behavior of quiescent LSCs in T-ALL.
Protocol
In this protocol, we are using GFP-labeled zebrafish T-ALL cells that were generated previously in the CG1 strain and thus can be directly injected into recipient syngeneic CG1 zebrafish15. Briefly, leukemia was generated by DNA microinjection of rag2:Myc and rag2:GFP into single-cell CG1 zebrafish embryos. Animals were monitored for leukemia development starting at 3 weeks post injection, using fluorescence microscopy. GFP-positive leukemia cells were FACS isolated and serially transplanted in recipient CG1 zebrafish to generate clones with a high frequency of LSC. Details of the entire protocol are described by Blackburn et al.15.
Alternatively, primary rag2:Myc-derived T-ALL can be generated by DNA microinjection of zebrafish embryos17. The DNA microinjection of rag2:Myc with a rag2-driven fluorescent reporter such as GFP can result in the development of B-cell, T-cell, and mixed ALL18. The leukemia clones used in this protocol were previously verified to be T-ALL15. All experimental procedures involving zebrafish were reviewed and approved by the University of Kentucky's Institutional Animal Care and Use Committee, protocol 2019-3399.
1. Propagating zebrafish T-ALL cells in vivo
- Thaw one vial containing 1 mL of frozen GFP-labeled zebrafish T-ALL cells in a 37 °C water bath with gentle shaking, until thawed. Once thawed, transfer to ice.
- Slowly pipette the contents of the tube into a 15 mL conical tube with 4 mL of fish media (0.9x PBS + 5% FBS) to dilute the freezing media.
- Spin down for 5 min at 2,500 × g at 4 °C. Remove media from the pellet. Resuspend in 0.5 mL of fish media and keep the cells on ice.
- Count the cells and dilute them if needed to 100,000 cells/5 µL.
- Anesthetize the adult CG1 zebrafish by adding 200 µL of 4 mg/mL anesthetic (MS-222) to 25 mL of fish system water in a Petri dish. Hold the CG1 fish, ventral side up and inject 5-6 µL into the intraperitoneal (IP) space using a Hamilton microliter syringe. Move the animals back to their tanks and monitor for a return to normal swimming before putting them back on the system.
NOTE: Adult zebrafish, 60 days or older were used for the transplantation studies described in this protocol. - Monitor leukemia growth once per week using a fluorescence microscope. See Section 2 below for details of the screening procedure.
2. Harvesting of fluorescently labeled leukemia cells
- Approximately 21-28 days post-transplant, anesthetize the adult CG1 zebrafish by adding 200 µL (4-5 drops) of 4 mg/mL anesthetic to 25 mL of fish system water in a Petri dish.
- Examine the CG1 zebrafish for leukemia burden by assessing the percentage of the GFP-labeled leukemia cells in the animal with an epifluorescent microscope, using the appropriate filter to detect the GFP fluorescence. The GFP-positive leukemia is ready to be harvested if it has spread across >70% of the animal.
- Euthanize the animal by adding 1 mL of 4 mg/mL anesthetic in a Petri dish containing 9 mL of fish system water. Allow all signs of life to cease; for instance, observe the fish for the cessation of opercular movement (~5 min).
- Move the zebrafish into a new Petri dish and add 1 mL of fish media to serve as a buffer for the cells. Using a new razor blade, first decapitate the zebrafish, then macerate the tissue using the same razor blade. Pipet up and down to dislodge large clumps of cells.
- Pass the cells through a 40 µm cell strainer into a 50 mL conical tube to dissociate into a single-cell suspension. Keep the cells on ice.
3. Optimization of the cell proliferation dye (CellTrace Far Red) concentration for staining and zebrafish cell viability
NOTE: For this protocol, and since the tumor cells are GFP-labeled, we used a far-red cell proliferation dye (CellTrace Far Red) to avoid spectral overlap, which will be referred to as CT-FR.
- Prepare a 1 mM solution of the fluorescent dye immediately before use by adding the appropriate volume of DMSO as per the manufacturer's recommendations. Have working solutions of 1 mM, 0.5 mM, 0.1 mM, and 0.01 mM; dilute as needed in fish media to test cell viability following staining.
- Count the cells and collect 1 × 106 cells in 1.5 mL microcentrifuge tubes.
- Centrifuge at 2,500 × g at 4 °C for 5 min. Remove the liquid and resuspend the cells in 1 mL of 0.9x PBS.
- Split the cell suspension between four tubes of 250 µL each for treatment with CT-FR.
- Add 250 µL of 0.9x PBS to bring the volume up to 500 µL each with approximately 250,000 cells/tube.
- Add the required amounts of the working solutions described above, to each of the four tubes to achieve the following final concentrations 1 µM, 0.5 µM, 0.1 µM, and 0.05 µM. Incubate for 20 min at 37 °C, protected from light.
- Pellet the cells by spinning down at 2,500 × g for 5 min at room temperature. Remove the liquid and wash the cells with 500 µL of fish media to remove the excess dye.
- Pellet again, remove the liquid, and resuspend in 25 µL of fish media. Use 1 µL to count cells in a 1:10 dilution in 50% Trypan Blue.
- Examine the cells under the microscope for cell viability; live cells will not retain the Trypan blue and appear bright under the microscope, while dead cells will take up the dye and appear dark. Examine the cells under the appropriate fluorescence channel first to identify tumor cells and then use the brightfield channel to assess cell viability.
- Select the concentration that results in successful staining while maintaining cell viability (>90% trypan blue-negative cells).
NOTE: For the zebrafish T-ALL cells tested in this experiment, 1 µM concentration was used for successful staining, and >90% of the cell viability.
4. Propagating CT-FR-stained T-ALL zebrafish cells in vivo
- Using the optimized CT-FR concentration from Section 3, stain the zebrafish cells. Leave some tumor cells unstained for the generation of the GFP-labeled T-ALL tumor, as a GFP-positive control during FACS sorting. Dilute both stained and unstained cells to 50,000 cells/5 µL for transplant.
- Anesthetize the adult CG1 zebrafish by adding 200 µL of 4 mg/mL anesthetic to 25 mL of fish system water in a Petri dish.
- Inject 5 µL of the cell suspension into the IP cavity of the zebrafish as described in Section 1 for both the stained and the unstained cells. Inject 5-6 zebrafish per group to account for the potential mortality post injection.
- Examine the animals by fluorescent microscopy under sedation 2x weekly for leukemia development.
NOTE: After ~10-14 days, the leukemia should occupy about 30-40% of this body. Plan for leukemia harvest and subsequent sort within this time frame. Waiting for the tumor to expand to a large size might result in a greater dilution of the fluorescent cell dye and difficulty in detecting its signal. Consider optimization of the timing of tumor harvest for the tumor model chosen. In addition, more frequent tumor monitoring is recommended here to ensure the capture of the desired tumor size.
5. Preparing for sorting of CT-FR-stained zebrafish T-ALL cells
- Prepare the following controls for the cell sorting: No color control (unstained, non-transgenic, single-cell suspension, from a wild type zebrafish), GFP-Labeled T-ALL cells, and newly stained CT-FR WT zebrafish cells (for setting a maximum threshold of Far-Red signal).
- Preparing unstained and CT-FR controls
- Harvest wild-type (WT) fish cells by euthanizing a non-transplanted CG1 zebrafish by adding 1 mL of 4 mg/mL anesthetic in a Petri dish containing 9 mL of fish system water.
- Move the zebrafish into a new Petri dish and add 1 mL of fish media to serve as a buffer for the cells. Use a new razor blade to first decapitate and then macerate the zebrafish using the same razor blade; pipet up and down to dislodge large cell clumps.
- Pass the cells through a 40 µm cell strainer into a 50 mL conical tube to dissociate into a single-cell suspension. Keep the cells on ice.
- For the CT-FR control, transfer 250,000 WT cells into a microcentrifuge tube and pellet the cells by centrifugation at 2,500 × g for 5 min at room temperature. Remove the fish media and add 500 µL of 0.9x PBS+ 1% FBS.
- Add 0.5 µL of the 1 mM CT-FR dye stock solution for a final concentration of 1 µM. Gently pipet with 1,000 µL pipet to mix. Incubate at 37 °C for 20 min, protected from light.
- Remove the liquid and wash the cells with 500 µL of fish media to remove the excess dye.
- Pellet the cells by centrifugation again, remove the liquid, and resuspend the cells in 500 µL of 0.9x PBS+ 1% FBS and pass through a 35 µm filter cap into a FACS tube. Keep on ice.
- Preparing the donor plate
NOTE: The donor plate is a 96-well v-bottom plate into which the CT-FR-stained T-ALL cells from transplanted zebrafish will be sorted. Each well of the plate contains 5,000 unstained, WT CG1 cells in 50 µL of fish media to serve as a carrier to help pellet the sorted cells in subsequent steps.- Count 500,000 WT CG1 cells and transfer them into a 15 mL conical tube.
- Dilute this cell stock to 100 cells/µL by adding 5 mL of fish media.
- Dispense 50 µL of the cell suspension into each well of the 96-well plate. Mix the cell suspension stock tube after each row to maintain cell suspension homogeneity. Keep the plate at 4 °C until sorting.
- Preparing the transplanted fish samples
- Select transplanted fish at 30-40% T-ALL burden. Collect fish transplanted with both the CT-FR-stained cells and unstained GFP tumor cells.
- Harvest cells as described in Section 2.
NOTE: Since the dye-retaining cells are expected to be present at a low abundance, prepare to sort a large number of cells from the CT-FR GFP sample. - Dilute into appropriate concentration for sorting ~3-5 × 106 cells/mL in 0.9x PBS + 1% FBS and pass through a 35 µm filter cap into a FACS tube. Add DAPI (1 mg/mL) at 1:1,000 dilution as a vital dye to exclude dead cells. Keep on ice.
- For the unstained GFP tumor cells, transfer 250,000 cells into a microcentrifuge tube and pellet at 2,500 × g for 5 min at room temperature. Remove the fish media, add 500 µL of PBS, and pass through a 35 µm filter cap into a FACS tube. Keep on ice.
6. Sorting
NOTE: Keep all tubes and the donor plate on ice the entire time except when used for sorting.
- For sorting parameters and gates, start by applying the forward and side scatter gating to identify the cells of interest based on their size and eliminate cell debris. Next, use forward scatter height versus forward scatter area to exclude duplicate cells and keep only singlets. Finally, use the no-color and single-color controls to draw the quadrants to identify GFP-positive and CT-FR-positive cell populations.
NOTE: The CT-FR control is expected to be much brighter than the samples for sorting since the staining for the control was done fresh with no cell divisions; cells from the CT-FR GFP samples are expected to appear at the lower end of the CT-FR positive gate. - Sort the number of cells to be transplanted into each fish into one well of the donor plate.
NOTE: For this experiment, we used a limiting dilution assay to estimate the frequency of leukemia stem cells in CT-FR High and CT-FR Low. This assay used a dose of 25 cells, transplanted into 3 animals, and a dose of 10 cells, transplanted into 10 animals. A typical limiting dilution assay using the zebrafish rag2:Myc-derived T-ALL would involve a group with a higher number of cells (500 or 1,000) since the LSC frequency in this model is estimated to be 1 LSC in hundreds of cells. However, in this assay, we are using one of the T-ALL clones previously described by Blackburn et al.15. The LSC frequency in the sample used here is ~1 LSC in tens of cells; we predict leukemia to be detected in animals transplanted in the 25 cells group. When optimizing this assay for your disease model, care should be taken while selecting the higher cell number group to ensure the engraftment of the tumor. - After sorting, keep the plate on ice until ready for transplant.
NOTE: Sorting procedures can be stressful and affect cell viability. Before running the actual limiting dilution experiment, a pilot sorting experiment was done. Sorted cells were transplanted and leukemia development was monitored to determine if the sorting was detrimental to cell viability. Alternatively, cell viability can be assessed by staining the sorted cells with Trypan blue and examining them under the microscope before transplant.
7. Transplanting sorted cells at limiting dilution and tracking tumor onset
- Spin down the plate using a large benchtop centrifuge at 2,500 × g at 4 °C.
- Carefully remove 45 µL of the supernatant from each well, leaving 5 µL liquid behind and making sure to avoid touching the cell pellet at the bottom of the well.
- Resuspend the cells with a 20 µL pipet. Resuspend 2-3 wells with the pipet and then using the Hamilton microliter syringe, inject the cell suspensions from each well into the desired number of CG1 zebrafish via IP transplant. Then, resuspend the cells in the next group of wells to avoid resettling of the cells.
- Examine the zebrafish by fluorescence microscopy under sedation once every week for T-ALL development for at least 6 weeks.
8. Determination of the leukemia stem cell frequency
- Record the number of positive animals per cell dose and treatment group at 6 weeks post transplant, considering animals with any amount of GFP signal to be positive. Consider animals to have significant leukemia burden when the GFP signal occupies >50% of the body.
- Use the web-based ELDA (Extreme Limiting Dilution Analysis) statistical software (http://bioinf.wehi.edu.au/software/elda/)19 to calculate the frequency of LSCs within each cell population.
NOTE: A schematic illustration of the workflow is provided in Figure 1.
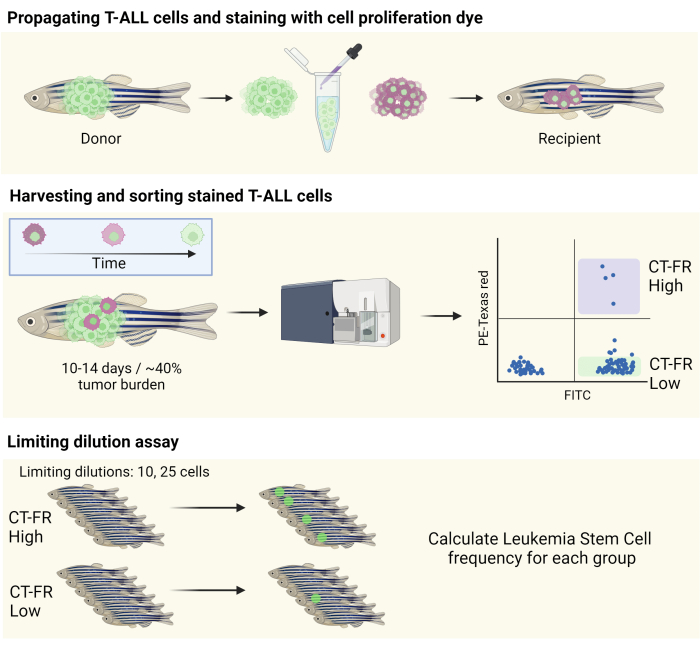
Figure 1: Workflow for the use of the cell tracking stain to isolate quiescent cells in the zebrafish T-ALL model. Schematic illustration of staining zebrafish T-ALL cells with the cell proliferation stain, propagation into a CG1 zebrafish (top panel), and sorting of cells based on the retention of the cell proliferation stain (middle panel). Collected cells were used for a limiting dilution assay to determine the frequency of LSCs (bottom panel). Abbreviations: T-ALL = T-cell acute lymphoblastic leukemia; LSCs = leukemic stem cells; CT-FR = CellTrace Far Red; FITC = fluorescein isothiocyanate; PE = phycoerythrin. Please click here to view a larger version of this figure.
Results
We followed the above-described protocol to sort cells that have retained the cell proliferation dye, CT-FR, and used them for a Limiting dilution assay (LDA) to estimate LSC frequency in the CT-FR High and CT-FR Low populations. To set the gating for the flow cytometry experiment, we used a no fluorophore (no color) control in addition to single-color controls (Figure 2A). Those quadrants were used to identify FITC- and PE-positive and -negative popula...
Discussion
LSCs are known to be resistant to conventional, anti-proliferative chemotherapy treatments, and finding targeted therapies against these cells holds great promise in reducing the occurrence of relapse and improving patient prognosis20. Previous research described the use of fluorescent cell proliferation stains to identify a small population of quiescent cells associated with drug resistance and stemness in T-ALL PDX models13. In this work, we describe the use of a similar ...
Disclosures
The authors have no conflicts of interest to disclose.
Acknowledgements
Funding for this research was provided by the National Cancer Institute (R37CA227656 to JSB). This research was also supported by the Flow Cytometry and Immune Monitoring Shared Resources of the University of Kentucky Markey Cancer Center (P30CA177558).
Materials
| Name | Company | Catalog Number | Comments |
| 26 G/2” micro-syringe | Hamilton | 87930 | NA |
| 35 µm filter cap FACS tubes | Falcon | 352235 | NA |
| 40 µm cell strainer | CELLTREAT | 229482 | NA |
| 96-well skirted PCR plate | Thermo Fisher Scientific | AB0800 | NA |
| Cell sorter | Sony Biotechnology | SY3200 | NA |
| CellTrace Far Red | Thermo Fisher Scientific | C34564 | NA |
| Conical tubes | VWR | 10026-078 | NA |
| DAPI | Thermo Fisher Scientific | 62248 | NA |
| DMSO | Sigma-Aldrich | D4818 | NA |
| Dulbecco'sPhosphate-buffered saline (PBS) | Caisson Labs | 22110001 | NA |
| Epifluorescence stereo microscope | Nikon | SMZ25 | NA |
| Fetal Bovine Serum (FBS) | Sigma-Aldrich | 12306C | NA |
| Fish system water | N/A | N/A | 0.03-0.05% salinity, pH 6.5-8, buffered with sodium bicarbonate |
| Microcentrifuge tubes | Thermo Fisher Scientific | C2171 | NA |
| MS-222 | Pentaire | TRS-1 | tricaine mesylate, an anesthetic |
| Petri dishes | Corning | 07-202-011 | NA |
| Razor blades | American Line | 66-0089 | NA |
| Trypan Blue | Thermo Fisher Scientific | T10282 | NA |
References
- Reya, T., Morrison, S. J., Clarke, M. F., Weissman, I. L. Stem cells, cancer, and cancer stem cells. Nature. 414 (6859), 105-111 (2001).
- Arai, F., et al. Tie2/angiopoietin-1 signaling regulates hematopoietic stem cell quiescence in the bone marrow niche. Cell. 118 (2), 149-161 (2004).
- Li, L., Bhatia, R. Stem cell quiescence. Clin. Cancer Res. 17 (15), 4936-4941 (2011).
- Kreso, A., Dick, J. E. Evolution of the cancer stem cell model. Cell Stem Cell. 14 (3), 275-291 (2014).
- Chen, W., Dong, J., Haiech, J., Kilhoffer, M. -. C., Zeniou, M. Cancer stem cell quiescence and plasticity as major challenges in cancer therapy. Stem Cells Int. 2016, 1740936 (2016).
- Kleffel, S., Schatton, T. Tumor dormancy and cancer stem cells: Two sides of the same coin. Adv Exp Med Biol. 734, 145-179 (2013).
- Tuy, K., Rickenbacker, L., Hjelmeland, A. B. Reactive oxygen species produced by altered tumor metabolism impacts cancer stem cell maintenance. Redox Biol. 44, 101953 (2021).
- Zhou, B. -. B. S., et al. Tumour-initiating cells: Challenges and opportunities for anticancer drug discovery. Nat Rev Drug Discov. 8 (10), 806-823 (2009).
- Lyons, A. B., Blake, S. J., Doherty, K. V. Flow cytometric analysis of cell division by dilution of cfse and related dyes. Curr Protoc Cytom. 64 (1), 11-12 (2013).
- Lyons, A. B. Analysing cell division in vivo and in vitro using flow cytometric measurement of cfse dye dilution. J Immunol Methods. 243 (1-2), 147-154 (2000).
- Azari, H., Deleyrolle, L. P., Reynolds, B. A. Using carboxy fluorescein succinimidyl ester (cfse) to identify quiescent glioblastoma stem-like cells. Methods Mol Biol. 1686, 59-67 (2018).
- Deleyrolle, L. P., Rohaus, M. R., Fortin, J. M., Reynolds, B. A., Azari, H. Identification and isolation of slow-dividing cells in human glioblastoma using carboxy fluorescein succinimidyl ester (cfse). J Vis Exp. (62), e3918 (2012).
- Ebinger, S., et al. Characterization of rare, dormant, and therapy-resistant cells in acute lymphoblastic leukemia. Cancer Cell. 30 (6), 849-862 (2016).
- Langenau, D. M., et al. Myc-induced t cell leukemia in transgenic zebrafish. Science. 299 (5608), 887-890 (2003).
- Blackburn, J. S., et al. Clonal evolution enhances leukemia-propagating cell frequency in t cell acute lymphoblastic leukemia through akt/mtorc1 pathway activation. Cancer Cell. 25 (3), 366-378 (2014).
- Al-Hamaly, M. A., Turner, L. T., Rivera-Martinez, A., Rodriguez, A., Blackburn, J. S. Zebrafish cancer avatars: A translational platform for analyzing tumor heterogeneity and predicting patient outcomes. Int J Mol Sci. 24 (3), 2288 (2023).
- Blackburn, J. S., Liu, S., Langenau, D. M. Quantifying the frequency of tumor-propagating cells using limiting dilution cell transplantation in syngeneic zebrafish. J Vis Exp. (53), e2790 (2011).
- Borga, C., et al. Simultaneous b and t cell acute lymphoblastic leukemias in zebrafish driven by transgenic myc: Implications for oncogenesis and lymphopoiesis. Leukemia. 33 (2), 333-347 (2019).
- Hu, Y., Smyth, G. K. Elda: Extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J Immunol Methods. 347 (1-2), 70-78 (2009).
- Bhojwani, D., Pui, C. -. H. Relapsed childhood acute lymphoblastic leukaemia. Lancet Oncol. 14 (6), e205-e217 (2013).
- Zon, L. I., Peterson, R. T. In vivo drug discovery in the zebrafish. Nat Rev Drug Discov. 4 (1), 35-44 (2005).
- Al-Hamaly, M. A., et al. Zebrafish drug screening identifies erlotinib as an inhibitor of wnt/β-catenin signaling and self-renewal in t-cell acute lymphoblastic leukemia. Biomed Pharmacother. 170, 116013 (2024).
- Yan, C., et al. Visualizing engrafted human cancer and therapy responses in immunodeficient zebrafish. Cell. 177 (7), 1903-1914 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved