A subscription to JoVE is required to view this content. Sign in or start your free trial.
Comprehensive Characterization of Tissue Mineralization in an Ex Vivo Model
In This Article
Summary
The proposed protocol entails a global approach to assess bone formation in the context of bone regeneration using multimodal analyses. It aims to provide qualitative and quantitative information on new bone formation, enhancing the rigor and validity of basic and pre-clinical investigations.
Abstract
The extensive characterization of tissue mineralization in the context of bone regeneration represents a significant challenge, given the numerous modalities that are currently available for analysis. Here, we propose a workflow for a comprehensive evaluation of new bone formation using a relevant large animal osseous ex vivo explant. A bone defect (diameter = 3.75 mm; depth = 5.0 mm) is created in an explanted sheep femoral head and injected with a macroporous bone substitute loaded with a pro-osteogenic growth factor (bone morphogenetic protein 2 - BMP2). Subsequently, the explant is maintained in culture for a 28-day period, allowing cellular colonization and subsequent bone formation. To evaluate the quality and structure of newly mineralized tissue, the following successive methods are set up: (i) Characterization and high-resolution 3D images of the entire explant using micro-CT, followed by deep learning image analyses to enhance the discrimination of mineralized tissues; (ii) Nano-indentation to determine the mechanical properties of the newly formed tissue; (iii) Histological examinations, such as Hematoxylin/Eosin/Saffron (HES), Goldner's trichrome, and Movat's pentachrome to provide a qualitative assessment of mineralized tissue, particularly with regard to the visualization of the osteoid barrier and the presence of bone cells; (iv) Back-scattering scanning electron microscopy (SEM) mapping with internal reference to quantify the degree of mineralization and provide detailed insights into surface morphology, mineral composition, and bone-biomaterial interface; (v) Raman spectroscopy to characterize the molecular composition of the mineralized matrix and to provide insights into the persistence of BMP2 within the cement through the detection of peptide bonds. This multimodal analysis will provide an effective assessment of newly formed bone and comprehensive qualitative and quantitative insights into mineralized tissues. Through the standardization of these protocols, we aim to facilitate interstudy comparisons and improve the validity and reliability of research findings.
Introduction
Bone defects, whether caused by trauma, tumor resection, congenital anomalies, or infection, represent a major challenge for regenerative medicine. These alterations compromise the structural integrity of the skeletal system, leading to discomfort, functional impairment, and a reduction in patients' quality of life.
To overcome these challenges, innovative bone repair strategies have emerged, with a focus on enhancing osteogenesis and bone tissue regeneration. These approaches include the use of implantable, injectable, or 3D-printable bone substitutes, which can be of natural origin (e.g., bio-sourced macromolecules, animal-derived hydroxyapatite) or synthetic (e.g., bioglasses, calcium phosphates)1. To enhance their low inherent ability to guide and stimulate bone regeneration, bone substitutes can be loaded with osteoinductive factors, such as bone morphogenetic proteins (BMPs), to promote osteogenic differentiation of progenitor cells and enhance bone formation2.
Bone formation is based on the initial formation of a collagen matrix, which is then mineralized by hydroxyapatite crystals, thereby reinforcing the bone structure3. This process confers specific stiffness and strength to the bone. The quality of the mineralized tissue is intricately governed by its microstructural attributes and degree of mineralization4. This quality plays a pivotal role in bone healing and the functionality of the regenerated bone5. However, characterizing bone mineralization remains a challenging task due to the inherent variability across multivariate studies6,7,8.
In addition, initial evaluations of the biocompatibility, cytocompatibility, and differentiation potential of bone graft substitutes are typically conducted in vitro. However, methodological disparities impede the seamless comparison of outcomes. Furthermore, these in vitro studies do not fully capture the multicellular interactions and complex dialogue between cell populations, including bone marrow cells, which are essential for regulating the bone regeneration process9. This lack of accurate representation of the bone microenvironment may compromise the accuracy of subsequent preclinical studies10.
Although in vivo assessments provide a more accurate representation of physiological contexts, they are constrained by ethical, logistical, and financial considerations. Consequently, ex vivo evaluations play a pivotal role as an interface between in vitro and in vivo studies, serving as a necessary intermediate step before moving on to experiments on living subjects11,12,13.
In this context, the implementation of comprehensive characterization methodologies is needed to appraise the quality of regenerated bone tissue and to ensure the relevance of the strategy before moving on to a preclinical model. Consequently, we propose a protocol based on the analysis of an explant model using sheep knee joint tissue. This innovative methodology involves implanting BMP2-loaded cement into the explants and conducting a detailed analysis of tissue mineralization after 28 days of culture.
The technical approaches employed in this study are diverse and complementary, collectively providing a comprehensive approach to evaluating the quality of regenerated bone tissue (Figure 1). High-resolution micro-CT imaging enables detailed 3D visualization of the bone structure, providing valuable insights into the mineral density, morphology, and integrity of the newly formed tissue. This technique is crucial for assessing the efficacy of bone regeneration and monitoring the progression of mineralization over time. Nanoindentation is a precise approach for determining the mechanical properties of the tissue, such as its hardness and strength. By measuring the response of the material to a force applied on a nanometric scale, this method enables the assessment of the robustness and quality of the mineralized tissue. Histological examinations using common staining such as hematoxylin/eosin/saffron (HES), Goldner's trichrome, and Movat's pentachrome provide invaluable insights into tissue structure and composition. These stainings' allow differentiation of the various tissue components, including cells, extracellular matrix, and mineral deposits, thereby enabling a comprehensive qualitative assessment of the bone regeneration process. Backscatter scanning electron microscopy (SEM) mapping offers a high-resolution visualization of the surface of the samples, allowing detailed analysis of the degree of mineralization of the bone matrix, as well as the interfaces between the implanted material and the host tissue. Finally, Raman spectroscopy provides information regarding the molecular composition of the tissue, particularly through the identification of specific components such as proteins, lipids, and minerals. This approach enables the characterization of the mineralized matrix and the detection of growth factors such as BMP2, thereby providing crucial information on the persistence of pro-osteogenic stimuli in the regeneration medium.
Using a multi-disciplinary approach, integrating various analytical techniques, our study aims to provide a thorough and comprehensive assessment of the quality of regenerated bone tissue, thus providing a solid basis for the evaluation of bone graft substitutes and their potential clinical application.
Protocol
This study has been approved by an ethics and animal welfare committee and by the French National Veterinary and Food Administration under number G44171.
1. Preparation and culture of osteochondral explants
- Harvest sheep joint specimens from freshly euthanized animals in an aseptic environment. Position the sheep in a supine position and shave the left hind limb. Prepare by disinfecting with alcohol around the knee joint. Use lateral parapatellar arthrotomy to expose the anterior and posterior cruciate ligaments, followed by patellar dislocation to reveal the femoral trochlea.
- Following tissue harvesting, create a 4.75 mm diameter defect in the lateral condyle using an osteochondral autograft transfer system.
- Using an orthopedic hammer, create a defect 10 mm in depth and retrieve the explant from the condyle. Subsequently, perform a second defect of 8 mm diameter, centered around the first, using an osteochondral autograft transfer system. This yields osteochondral explants measuring 8 mm x 10 mm, containing a central defect of 4.75 mm x 10.0 mm.
- If required for logistical reasons, transport the explants in Hank's Balanced Salt Solution (hBSS) supplemented with 1% penicillin-streptomycin and store at 4 °C.
- Rinse the explants 3x with Phosphate Buffered Saline (PBS), then inject the 4.75 mm x 10.0 mm defect with a calcium phosphate cement supplemented with 40 µg/mL of BMP2. Prepare the cement as described below.
- Produce α-tricalcium phosphate (α-TCP) by heat treatment of apatitic tricalcium phosphate rods (isostatic press at 120 MPa) at 1364 °C for a minimum of 12 h, followed by air-quenching14.
- Grind the α-TCP rods in absolute ethanol with a planetary mill ball 2x for 5 min at 500 rpm to produce a fine powder of 13.1 ± 1.7 µm (measured using laser diffraction, as previously described14).
- Sterilize the α-TCP powder by dry heating it at 180 °C for 45 min. Prepare the calcium phosphate cement by mixing α-TCP powder with a 2.5% (w/v) Na2HPO4 0.22 µm filtered solution (liquid/powder ratio of 0.35) for 1 min.
- Load the cement paste into a 3 mL syringe prior to injection using an 18 G needle.
- After 10 min at room temperature (RT), transfer the explants into a 25 cm² flask and gently add 10 mL of complete culture medium consisting of Dulbecco's Modified Eagle Medium (DMEM) high glucose supplemented with 10% fetal bovine serum and 1% penicillin-streptomycin.
- Maintain the explants for up to 28 days in a 37 °C incubator with 5% CO2 and change the complete culture medium every 2 days under a laminar flow hood.
- Rinse the explants 3x with PBS. Transfer them into 50 mL tubes, add 20 mL of 4% paraformaldehyde (PFA), and keep them at 4 °C for 3 days. Then, rinse the explants with PBS and store them in 70% ethanol before subsequent analyses.
2. Micro-CT analysis
- Conduct micro-tomography analyses utilizing a micro-CT X-ray. Acquire X-ray projections at 10.7 µm resolution, with an exposure time of 1200 ms and a 1 mm aluminum filter (80 kV and 125 µA). Average three images for each 0.45° rotation increment in order to enhance the signal-to-noise ratio.
- Reconstruct three-dimensional (3D) images using the manufacturer's reconstruction software after an X/Y alignment with a reference scan, with the following parameters - Smoothing: 0, Ring Artefact: 3, Beam Hardening: 35%. Clean the image stacks with image processing software and observe with two-dimensional (2D) and 3D visualization tools.
3. Deep-learning image analyzes
- Conduct image segmentation using dedicated specialized software. Use the integrated segmentation Wizard to train a deep-learning model for bone and cement discrimination.
- Select a representative zone (i.e., frame) containing bone, cement, and background from the reconstructed micro-CT images. Segment this first frame manually using either simple (e.g., threshold) or powerful tools (e.g., propagation) provided by the software .
- Generate a deep learning model in the Model tab and select 3D-Unet Routine in the drop-down menu. Define its experimental parameters in adequation with the images to analyze by a right click on the generated model; here: depth of 5, patch size 32 x 32, Adadetla algorithm, stride ratio of 0.25, data augmentation x10.
- Use the segmented frame to train the deep-learning routine (click on the Train Button), which is acknowledged to be relevant for its fast and accurate image segmentation capabilities3,15. Once the training is completed, define a second work zone (frame) and automatically segment using the predict function.
- Apply manual correction if required to generate more accurate training data. Train routine once again. Repeat the process until a satisfactory result is obtained; the number of data augmentations progressively decreases as the number of training frames increases.
NOTE: The DICE index provided during the model training gives an indication of the accuracy of the model (compared to the input information) but is not sufficient. Confirmation of the relevance of the model should be carried out by validation of the automatically segmented data by 3 independent seasoned researchers.
- Apply manual correction if required to generate more accurate training data. Train routine once again. Repeat the process until a satisfactory result is obtained; the number of data augmentations progressively decreases as the number of training frames increases.
- Publish the model by clicking on the Export button and apply it to the entire micro -CT dataset by clicking Segment > Exported Model > Segment Full Dataset.
4. Embedding
- Dehydrate the bone explants by placing them in a 40 mL glass vial containing 25 mL of dehydration solution composed of 70% acetone and 30% xylene. Place the glass vial on a rotating wheel for 1 h at RT. Repeat this step 3x.
- Replace the dehydration solution with 25 mL of xylene and place the glass vial on the rotating wheel for 1 h at RT.
- Replace the xylene solution with 25 mL of methylmethacrylate enriched (MMA) with 10% benzoyl peroxide (BPO) and 10% dibutyl phthalate (DBP). Place it on the rotating wheel for 1 h at RT. Repeat this step 2x.
- Replace the MMA solution with 25 mL of MMA enriched with 10% BPO, 10% DBP, and 450 µL of N, N dimethylaniline diluted 1:20 in propan-2-ol. Place the glass vial at -20 °C overnight. The solution becomes yellowish.
- Place the explant into a medium-size embedding mold and pour the MMA-BPO-DBP-N, N aniline into the mold. Place the mold into a plastic box and ventilate with nitrogen flow for 5 min. Close the box hermetically and place it at 4 °C for 48 h for MMA polymerization and hardening.
- Remove the resin containing the explant from the mold and keep it at 4 °C until processed for subsequent analysis.
5. Scanning electron microscopy (SEM) - quantitative backscattered electron imaging (qBEI)
- Cut the pMMA block containing the explant with a diamond saw along its long axis. Keep the first half of the block for histological analysis. Cut the other half further to generate a 1.5 mm thick section. Perform the cut at 3000 rpm with a 3 mm/min speed.
- Grind the section with silicon carbide paper with ascending numbers corresponding to lower grain size: SiC 320 for 10 s, SiC 1000 for 15 s, SiC 2000 for 30 s, and SiC 4000 for 30 s.
- Polish the section with diamond paste and specific clothes: polishing cloth with 3 µm Mol B3 diamond solution for 1 min and fleece cloth with 1 µm Nap B1 diamond solution for 1 min.
- Rinse the section under double distilled water and clean off diamond particles with a cotton bud. Dry the section under nitrogen gas.
- Carbon coat the thick section (10 nm carbon film thickness) and mount it on an aluminum stub with silver paint. Make a silver paint bridge between the section top and the stub to allow for electron charging evacuation to the ground.
- Place the stub on the SEM stage. Next to the sample, insert a control stub composed of a Faraday cup with carbon, aluminum, and silicon standards into the SEM chamber. This will be used to calibrate the electron beam and convert the grey levels into percentages of Ca. Close the SEM chamber and vacuum.
- Turn on the electron beam and adjust SEM parameters to run in the backscattered electron mode. On backscattered images, the grey level of the carbon standard is 25, the aluminum standard is 225, and the silicon standard is 253. As an indication, on the system used, SEM parameters are accelerating voltage 20 keV, probe current measured with the Faraday cup: 250 pA, working distance: 15 mm, aperture: 30 µm, image resolution: 1024 x 768 pixels, vacuum: < 4.10-4 Pa, dwell time of 100 µs/pixel, brightness ~38 and contrast ~72.
- When the SEM is calibrated with the standards, acquire images of the specimen in backscattered electron mode.
- Use the backscattered electron image of the specimen to convert the grey levels to calcium content, as described in Roschger et al.16. Perform this using any image analysis software.
- Plot the distribution of calcium content. Calcium content distribution shows a Gaussian distribution. Three main parameters of interest are computed: Camean, as the mean calcium content on the image, Capeak as the most frequent calcium concentration encountered on the image and Cawidth as the full width at half maximum of the calcium content distribution.
NOTE: The reproducibility of qBEi has been investigated by imaging the same region of interest for 5 consecutive days (one acquisition per day). This means that the electron beam was turned on and off between sessions. The error percentage in Camean, Capeak, and Cawidth were estimated as 0.5%, 0.7%, and 1.2%, respectively. Calcium content was also investigated by Energy dispersive X-ray analysis (EDS). A very good linear relationship was established between Camean and calcium content estimated by EDS with an R² value of 0.99, similar to data obtained by Roschger et al. previously17.
6. Histology
- For each sample, cut 7 µm sections using a hard tissue microtome with a tungsten blade 18,19. Stain sections with Goldner trichrome, HES, and MOVAT stain using an automatic staining system.
- For deplastification, soak the samples successively in acetone for 5 min and repeat 2x. Rinse the samples in distilled water for 5 min and repeat 2x.
- Perform Goldner Trichrome coloration as described below.
- Place samples in Weigert ferric hematoxylin for 20 min. Wash with tap water. Immerse samples in 0.5% acid alcohol for 30 s-1 min. Wash with tap water for 20 min.
- Place samples in Ponceau/fuchsine acid/azophloxin solution for 5 min. Immerse in 1% acetic acid for 1 min.
- Stain with phosphomolybdic acid/Orange G solution for 20 min. Immerse in 1% acetic acid for 1 min.
- Stain with fast green solution for 15 min at room temperature or 8 min at 60 °C. Rinse thoroughly with tap water; repeat 3x.
- Perform Movat Pentachrome coloration as described below.
- Alcian Blue staining: immerse samples in Alcian Blue solution for 30 min. Rinse samples in tap water for 5 min. Place samples in alkaline ethanol for 60 min. Rinse samples in tap water for 10 min. Perform a final rinse with distilled water.
- Weigert Hematoxylin staining: place samples in Weigert's Ferric Hematoxylin for 20-30 min. Rinse samples in tap water for 15 min. Rinse again with distilled water.
- Staining with Crocein Brilliant/fuchsine acid: immerse the samples in a solution of Crocein Brilliant/Acid Fuchsin for 10 min. Rinse with 0.5% acetic acid. Immerse samples in a 5% phosphotungstic acid solution for 20 min. Rinse with 0.5% acetic acid for 2 min. Perform three successive rinses with 100% ethanol, each lasting 5 min.
- Staining with Safran du Gatinais: place samples in Safran du Gatinais for 15 min. Perform a final rinse with tap water.
- Perform HES coloration as described below.
- Staining with Weigert's Hematoxylin: submerge the samples in Weigert's Hematoxylin for 30 min. Rinse with tap water for 2 min.
- Decolorization: dip the samples in hydrochloric acid alcohol for 10 s. Rinse with tap water for 2 min.
- Neutralization: immerse the samples in a solution of lithium carbonate for 1 min. Rinse with tap water for 2 min. Perform a final rinse with distilled water for 1 min.
- Staining with Eosin-Erythrosine: place the samples in a solution of Eosin-Erythrosine for 3 min. Rinse with tap water for 10 s.
- Dehydration: Submerge the samples in 95% alcohol for 15 s. Transfer to 100% alcohol for 15 s. Repeat the immersion in 100% alcohol for 30 s.
- Final staining with Alcoholic Safranin: immerse the samples in alcoholic safranin for 10 min. Perform a final rinse with tap water.
- Perform slide mounting as described below.
- Rinse samples in 95% ethanol, then 100% ethanol, then methylcyclohexane and repeat this 3x for each step.
- Mount the samples using a mounting kit and add a coverslip. Scan the images using a digital slide scanner.
7. Raman microspectroscopy
- Use the same section used for Raman analysis as the SEM. Briefly grind and polish the section as described in steps 5.2 - 5.3 to remove the thin carbon layer added for SEM that impairs heat dissipation from the sample.
- Place the section on the stage of the Raman microspectrometer. Calibrate the wavenumber to ensure accuracy and align the laser prior to measurement. As such procedures are specific to each instrument, they will not be described in this article.
- Locate regions of interest in the tissue that need to be analyzed. Position the zone to be analyzed in the center of the video tool and collect Raman spectra with the following parameters: 785 nm laser used at 30 mW, time of integration: 20 s repeated 3 times, spectral range: 350-1800 cm-1, grating 1200 lines/mm.
- Collect 10 spectra of the embedding resin with the same Raman settings and average them. Use this to subtract the resin contribution in step 7.6.
- Pre-process the spectra as follows to remove the contribution of fluorescence, noise, and resin: baseline correction with a five-order polynomial fitting, Savitzky-Golay filter with a degree of 4, and a window size comprised between 17 and 21, digital resin subtraction using the ~812 cm-1 peak of the resin. Such pre-processing can be performed with different software designed for data analysis and visualization.
- Further analyze the pre-processed spectra to extract valuable parameters by rationing the peak intensity of the vibration of interest with the peak intensity of the reference vibration. For bone analysis, for example, use the following parameters to compute it.
- Mineral-to-matrix ratios: divide the peak intensity of the v1PO4 vibrations at ~960 cm-1 by the peak intensity of the Amide I (~1668 cm-1), amide III (~1250 cm-1), CH2-wag (~1450 cm-1) or the sum of proline (~854 cm-1) and hydroxyproline (Hyp, ~872 cm-1) vibrational modes. The v2PO4, located at ~430 cm-1, or the v4PO4, located at ~600 cm-1, can also be used and ratioed with the Amide III vibration20. These parameters represent the degree of mineralization of the organic phase of the bone matrix.
- Carbonate/Phosphate: divide the peak intensity of the v1CO3, located at ~1070 cm-1, by the v1PO4. This represents the amount of type B carbonate substitution in the apatite lattice. Hyp/Pro, represents the hydroxyproline content, ~1375 cm-1/Amide III, represents the proteoglycan content, ~1150 cm-1/CH2-wag and ~1495 cm-1/CH2-wag, represents the amount of the advanced glycation end products carboxymethylysine and pentosidine, 1670 cm-1/1690 cm-1 and 1670 cm-1/1640 cm-1 represent the disordered secondary structure of the collagen, also known as collagen maturity, and ordered structures in the form of α-helix in type I collagen, respectively.
- Evaluate the reproducibility of Raman measurements by examining the same region of interest in 5 consecutive acquisitions (one acquisition a day). For each analyzed parameter, the variance was below 0.2% of the parameter value, supporting good reproducibility. The specificity of each investigated peak has been extensively discussed in the field, and an overview is presented in the recent review of Unal21.
8. Nanoindentation
NOTE: Due to the destructive nature of nanoindentation, it is usually performed at the end of the sample analysis routine. The nanoindentation system that we own is equipped with a pyramidal Berkovitch diamond indenter. However, several indenter shapes exist, and no consensus in the literature has been determined for bone or biomaterial specimens.
- Hydrate the embedded bone specimen in saline for 16 h at RT prior to nanoindentation testing.
- Calibrate the nanoindentation system using fused silica and record the resulting indentation modulus. For fused silica, the indentation modulus is approximately 72 GPa using a Poisson coefficient of 0.17.
- Once the nanoindentation system is calibrated, place the sample used for Raman analyses in the optical system of the nanoindentation device to point out locations where nanoindentation will be performed.
NOTE: This modality varies from one equipment to another; we do not detail how the nanoindentation system and software work but give general advice about the different steps that should be performed. - Once the location of nanoindentation is chosen, move the sample under the indentation device and perform indentation. Perform nanoindentation at a constant depth of 900 nm, with a loading/unloading speed of 40 mm/min and 15 s pause between loading and unloading. Set the Poisson coefficient for bone tissue at 0.3.
- Retrieve the different parameters from the nanoindentation curves and compute them according to Oliver and Pharr's theory22.
- Indentation modulus (EIT) is derived from the known properties of the indentation tip and the reduced modulus and combines the local elastic modulus of the specimen. Compute indentation as:
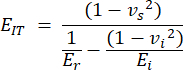
With vs, vi, Ei, and Er representing the Poisson's coefficient of the sample, assumed to be 0.3 for bone, the Poisson's coefficient of the diamond indenter, assumed to be 0.07, the elastic modulus of the diamond indenter, fixed at 1140 GPa and the reduced modulus, computed as:
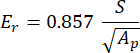
With S representing the slope of the unloading segment and Ap being the projected area and computed as:
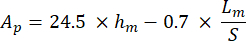
- Hardness (HIT): corresponding to the bone resistance to initiation and propagation of cracks. In other words, it is an indicator of bone toughness. Compute HIT as:
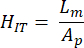
- Indentation creep rate (CIT): This reflects the bone deformation over time at constant load, indicative of the visco-elastic properties of the specimen, calculate as:
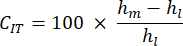
- Maximum load (Lm): This corresponds to the opposed loading necessary to penetrate the indicated depth into the mineralized bone matrix.
- Indentation work (Wplast): This corresponds to the area under the load/unload deformation curve. It is a direct reflection of the energy dissipated to induce plastic deformation.
NOTE: The reproducibility of nanoindentation is more difficult to assess on bone samples due to the destructive nature of this methodology. Nevertheless, we estimated the reproducibility of the standard and uniform fused silica sample. For all of the above parameters, the variance was estimated below 0.26% (5 consecutive measures on 5 separate days), supporting very good reproducibility. On the other hand, due to the anisotropic nature of bone samples, it is more difficult to appreciate their reproducibility.
- Indentation modulus (EIT) is derived from the known properties of the indentation tip and the reduced modulus and combines the local elastic modulus of the specimen. Compute indentation as:
Results
A micro-CT image of the explant is shown in Figure 2. Using Manual segmentation cannot optimally separate bone from cement, present in the central canal, using global thresholding. To improve the recognition of trabecular bone and cement, we propose to use deep learning. Deep learning is powerful for recognizing biomaterial characteristics and helps to improve the separation between bone and cement, enabling a better assessment of cement-bone interactions. This is of the utmost importance in...
Discussion
Repair of bone defects is a major challenge in regenerative medicine to restore mobility, reduce pain, and improve the quality of life of affected individuals. The use of explant models offers a number of advantages compared to in vivo studies for the investigation of bone defect repair. In addition to ethical considerations, this model allows for the rigorous control of experimental conditions and the reduction of biological variability, thereby facilitating the generation of more accurate and reproducible resu...
Disclosures
The authors have no conflicts of interest to disclose.
Acknowledgements
We want to thank the technical facilities involved in the collection and processing of specimens, including SC3M (SFR Francois Bonamy (UMS 016), University of Nantes), SFR ICAT (University of Angers), BIO3, HiMolA, and SC4BIO. The Inserm UMR_S 1229 RMeS is supported by grants from the French Government through Inserm, Nantes Université, Univ Angers and Oniris VetAgroBio institutions. CL is also grateful to HTL Biotechnology.
Materials
| Name | Company | Catalog Number | Comments |
| 0.20 filters | VWR | 28145-501 | |
| 18 G needle (1,2x40 mm) | Sterican | 4665120 | |
| 3 mL syringe | HENKE-JECT | 8300005762 | |
| 37% hydrochloric acid | VWR | 1.00317.1000 | |
| Acetic acid (glacial) | Sigma | A6283 | |
| Acetone | VWR | 20063-365 | |
| Alcian Blue 8GX | VWR | 361186 | |
| Ammonium hydroxide | VWR | 318612 | |
| Apatitic tricalcium phosphate | Centre for Biomedical and Healthcare Engineering (Mines Saint Etienne, France) | TV26U | |
| Azophloxine | Sigma | 210633 | |
| Benzoyl peroxide | Sigma | 8.01641.0250 | |
| BMP2 | Medtronic | InductOs 1.5 mg/mL | |
| Brillant crocein | Aldrich | 2107507 | |
| CTVox | Bruker | - | |
| DataViewer | Skyscan | - | |
| Diamond blade | Struers | MOD13 | |
| Diamond saw | Struers | Accutom-50 | |
| DiaPro Mol B3 diamond solution | Struers | 40600379 | |
| DiaPro Nap B1 diamond solution | Struers | 40600373 | |
| Dibasic sodium phosphate (Na2HPO4) | Sigma | 102404598 | |
| Dibutyl Phtalate | Chimie-Plus Laboratoires | 28656 | |
| DragonFly software | ORS | 2022.1.0.1231. | |
| Dulbecco's Modified Eagle Medium (DMEM) high glucose, GlutaMAX(TM), pyruvate | ThermoFisher Scientific | 31966-021 | |
| Eosine Y- Surgipath | Sigma | 1002830105 | |
| Erythrosin B | Sigma | 102141057 | |
| Ethanol absolute | VWR | 20820362 | |
| Eukitt | Dutscher | 6.00.01.0003.06.01.01 | |
| Falcon 50 mL | Sarstedt | 62.547.254 | |
| Ferric chloride hexahydrate (FeCl3, 6H2O) | Merck | 1.03943.0250 | |
| Fetal Bovine Serum (FBS) | Eurobio | CVFSVF00 | |
| Fuchsine acid | Merck | 1.05231.0025 | |
| Hank's Balanced Salt Solution (HBSS) | Biosera | MS01NG100J | |
| Hematoxylin | Sigma | 86.118.9 | |
| Isostatic press | Nova Suisse | Pmax 1500 bars | |
| Laser diffraction granulometry | Malvern | Mastersizer 3000 | |
| Light green | Prolabo | 28947135 | |
| Lithium carbonate | Sigma | A13149 | |
| MD-Mol polishing cloth | Struers | 40500077 | |
| Methylcyclohexane | VWR | 8.06147.1000 | |
| Methylcyclohexane | VWR | 8.06147.1000 | |
| Methylcyclohexane | VWR | 8.06147.1000 | |
| Methylmethacrylate | Sigma | 8.00590.2500 | |
| Micro-CT, micro-scanner | Bruker | Skyscan 1272 | |
| Monobasic sodium phosphate (NAH2PO4) | Sigma | 71496 | |
| Mortar | Fritsch | Pulverisette 6 | |
| N,N, Dimethylanilin | Sigma | 803060 | |
| Nanoindentation station | Anton Paar | NHT2 | |
| ND-Nap polishing cloth | Struers | 40500080 | |
| OATS Osteochondral Autograft Transfer System Set, 4,75 mm | Arthrex | AR-1981-04S | |
| OATS Osteochondral Autograft Transfer System Set, 8 mm | Arthrex | AR-1981-08S | |
| Orange G | Ral | M15 | |
| Paraformaldehyde (PFA) | Sigma | P6148 | |
| Peel-a-way disposable embbedding moulds | Polysciences, Inc | 18646C-1 | |
| Penicillin/Streptomycin (P/S) | ThermoFisher Scientific | 15140122 | |
| Phosphate Buffered Saline (PBS) | ThermoFisher Scientific | 10010023 | |
| Phosphomolybdic acid | Sigma | 221856-100 g | |
| Phosphotungstic acid | Aldrich | 12863-5 | |
| Polishing machine | Sturers | Dap V | |
| Poupinel | MEMMERT | TV26U | |
| Raman microspectrometer | Renishaw | InVia Qontor | |
| Safran du Gâtinais | Labonord | 11507737 | |
| Scanning electron microscope | Carl Zeiss | Evo LS 10 | |
| SEM | Zeiss | Carl Zeiss Evo LS10 | |
| SiC foils/Grinding papers | Struers | 40400008 (#320), 40400011 (#1000), 40400122 (#2000), 40400182 (#4000) | |
| Silver paint | Electron microscopy sciences | 12686-15 | |
| Standard stub with Faraday cup, carbon, aluminium and silicon standards | Micro-Analysis Consultants Ltd | 8602 | |
| T25 flask | Corning | 430639 | |
| Xylene | VWR | 28975.325 | |
| Xylidine Ponceau | Aldrich | 19.976-1 |
References
- Feroz, S., Cathro, P., Ivanovski, S., Muhammad, N. Biomimetic bone grafts and substitutes: A review of recent advancements and applications. Biomed Eng Adv. 6, 100107 (2023).
- Tsuji, K., et al. BMP2 activity, although dispensable for bone formation, is required for the initiation of fracture healing. Nat Genet. 38 (12), 1424-1429 (2006).
- Hadjidakis, D. J., Androulakis, I. I. Boneremodeling. Ann N Y Acad Sci. 1092, 385-396 (2006).
- Boivin, G., Meunier, P. J. The degree of mineralization of bone tissue measured by computerized quantitative contact microradiography. Calcified Tiss Int. 70 (6), 503-511 (2002).
- Boivin, G., et al. Influence of remodeling on the mineralization of bone tissue. Osteoporo Int. 20 (6), 1023-1026 (2009).
- Zanghellini, B., et al. Multimodal analysis and comparison of stoichiometric and structural characteristics of parosteal and conventional osteosarcoma with massive sclerosis in human bone. J Str Biol. 216 (3), 108106 (2024).
- Trento, G., et al. formation around two titanium implant surfaces placed in bone defects with and without a bone substitute material: A histological, histomorphometric, and micro-computed tomography evaluation. Clin Implant Dent Relat Res. 22 (2), 177-185 (2020).
- Palmquist, A. A multiscale analytical approach to evaluate osseointegration. J Mater Sci Mater Med. 29 (5), 60 (2018).
- Budán, F., et al. Novel radiomics evaluation of bone formation utilizing multimodal (SPECT/X-ray CT) in vivo imaging. PLoS ONE. 13 (9), e0204423 (2018).
- Hulsart-Billström, G., et al. A surprisingly poor correlation between in vitro and in vivo testing of biomaterials for bone regeneration: results of a multicentre analysis. Eur Cells Mater. 31, 312-322 (2016).
- Cramer, E. E. A., Ito, K., Hofmann, S. Ex vivo bone models and their potential in preclinical evaluation. Curr Osteoporo Rep. 19 (1), 75-87 (2021).
- Chan, M. E., et al. A trabecular bone explant model of osteocyte-osteoblast co-culture for bone mechanobiology. Cell Mol Bioeng. 2 (3), 405-415 (2009).
- Doke, S. K., Dhawale, S. C. Alternatives to animal testing: A review. Saudi Pharma J. 23 (3), 223-229 (2015).
- Moussi, H., et al. New formulation of injectable and degradable calcium phosphate/silanized hyaluronic acid composite foam: Investigation in a rabbit model of long bone defect. SSRN. , (2024).
- Paré, A., et al. Standardized and axially vascularized calcium phosphate-based implants for segmental mandibular defects: A promising proof of concept. Acta Biomater. 154, 626-640 (2022).
- Roschger, P., Fratzl, P., Eschberger, J., Klaushofer, K. Validation of quantitative backscattered electron imaging for the measurement of mineral density distribution in human bone biopsies. Bone. 23 (4), 319-326 (1998).
- Roschger, P., Plenk, H., Klaushofer, K., Eschberger, J. A new scanning electron microscopy approach to the quantification of bone mineral distribution: backscattered electron image grey-levels correlated to calcium K alpha-line intensities. Scanning Micro. 9 (1), 75-86 (1995).
- Porter, A., et al. Quick and inexpensive paraffin-embedding method for dynamic bone formation analyses. Sci Rep. 7, 42505 (2017).
- Goldschlager, T., Abdelkader, A., Kerr, J., Boundy, I., Jenkin, G. Undecalcified bone preparation for histology, histomorphometry and fluorochrome analysis. J Vis Exp. (35), e1707 (2010).
- Kazanci, M., Roschger, P., Paschalis, E. P., Klaushofer, K., Fratzl, P. Bone osteonal tissues by Raman spectral mapping: Orientation-composition. J Str Biol. 156 (3), 489-496 (2006).
- Unal, M., Ahmed, R., Mahadevan-Jansen, A., Nyman, J. S. Compositional assessment of bone by Raman spectroscopy. Analyst. 146 (24), 7464-7490 (2021).
- Oliver, W. C., Pharr, G. M. An improved technique for determining hardness and elastic modulus using load and displacement sensing indentation experiments. J Mater Res. 7 (6), 1564-1583 (1992).
- Proffen, B. L., McElfresh, M., Fleming, B. C., Murray, M. M. A comparative anatomical study of the human knee and six animal species. Knee. 19 (4), 493-499 (2012).
- Allen, M. J., Houlton, J. E. F., Adams, S. B., Rushton, N. The surgical anatomy of the stifle joint in Sheep. Vet Surg. 27 (6), 596-605 (1998).
- Kleuskens, M. W. A., van Donkelaar, C. C., Kock, L. M., Janssen, R. P. A., Ito, K. An ex vivo human osteochondral culture model. J Ortho Res. 39 (4), 871-879 (2021).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved