A subscription to JoVE is required to view this content. Sign in or start your free trial.
A Xenograft Mouse Model to Assess Efficacy of Therapeutic Agents for Human Acute Leukemia
Not Published
* These authors contributed equally
In This Article
Summary
Mouse (Mus Musculus) models are being widely used to develop xenografts using human leukemia cells. These models provide a comparable biological system to study drug efficacy, pharmacodynamics, and pharmacokinetics. Modeling acute myeloid leukemia in immunocompromised mice is described in detail using the U937 cell line xenograft as an example.
Abstract
Preclinical evaluation of therapeutic agents using an appropriate animal model is a critical step and a requirement for selecting drugs worth testing in humans. Therapeutic agents such as small molecule inhibitors, biological agents, immune checkpoint inhibitors, and immunotherapy each have unique mechanisms of action and call for careful selection of in vivo systems in which their efficacy can be tested. The purpose of this article is to describe in detail development of one such leukemia xenograft model for testing the therapeutic efficacy of novel agents. Using an immunocompromised (NRG) murine model that lacks B, T, and NK cells helps engraftment of transplanted leukemia cells and provides an acceptable microenvironment to study the therapeutic efficacy of small molecule inhibitors and some biological agents. This article describes the development of leukemia murine xenografts for in vivo drug testing using an acute myeloid leukemia (AML) cell line murine model treated with the cytotoxic drugs daunorubicin and cytarabine as an example. Treatment response can be assessed during therapy using several noninvasive and minimally invasive methods. Bioluminescence imaging can be used to measure leukemia burden over time when luciferase prelabeled leukemia cells are used to develop xenografts. Peripheral blood count analysis provides vital information about side effects such as myelosuppression (e.g., cytopenia) and therapeutic effect (e.g., blast count or differentiation). These techniques help track differences in the development of leukemia or decrease in tumor burden at various time points during the drug treatment without scarifying the study animals. Secondary methods such as immunophenotyping using flow cytometry are applied to confirm differences in the leukemia burden among treated and untreated groups. The methods described here can be tailored and used for developing xenografts of other types of leukemia (e.g., acute lymphoblastic leukemia).
Introduction
Acute myeloid leukemia (AML) is a clonal disorder arising from a malignantly transformed multipotent hematopoietic stem cell that acquires consecutive genomic alterations, eventually advancing into clinically overt disease. It is a highly complex disease with significant genetic, epigenetic, and phenotypic heterogeneity1. The uncontrolled proliferation and impaired differentiation of myeloid precursor cells (i.e., blasts) is one of the hallmarks of AML, leading to anemia, thrombocytopenia, and eventually death2. According to the American Cancer Society, in 2019, ~21,450 new cases of AML will be diagnosed, and ~10,920 people will succumb to the disease3. Standard therapeutic options include cytarabine-based chemotherapy and hematopoietic stem cell transplantation (HSCT). The 5-year overall survival (OS) of patients younger than 60 years old is around 40%, and for those older than 60 years it is only 10−20%4.
Novel drug discovery and drug development is a formidable challenge for the scientific community as well as the pharmaceutical industry. On average, the development of a novel drug costs ~$2.6 billion and takes over 10 years5. Drug discovery for anticancer drugs is an inefficient and cumbersome process with 89% of drugs failing in preclinical testing to gain FDA approval6. Flawed preclinical research is one of the reasons for drug failures7. Although multiple models of cell culture and in vitro studies are useful and important for testing potential therapies, the drawback of cell line models is that the synthetic nature of their culture conditions means they do not necessarily reflect the behavior of the original cancer cells in patients8. Also, it is impossible to fully recapitulate the complexity of the whole organism and the bone marrow microenvironment in cell culture8. As a result, cell line-derived xenograft tumor mouse models were generated through the transplantation of well-established cancer cell lines into immunocompromised mice. The major advantage of the cell line-derived xenograft mouse models is that they more closely simulate the tumor’s microenvironment and pathophysiological conditions9.
Here, we provide a comprehensive protocol explaining the steps to generate leukemia xenograft mouse models using a stable AML cell line labeled with a luciferase reporter (U937-Luc-tdTomato). Details of lentiviral transduction of the leukemia cell line will not be explained10,11. We also describe a detailed protocol to monitor leukemia progression using bioluminescence imaging (BLI). Statistical analysis and reporting of the differences observed in leukemia progression in mice treated with control, with a single standard of care drug (i.e., cytarabine or daunorubicin), or a combination of both, will be detailed. The first part of this protocol (sections 1, 2, and 3) are devoted to selection of the mouse strain and generation of the xenograft model, where we describe the transplantation procedure followed by imaging, randomization, and drug treatment of mice. Later, we detail steps to collect leukemia cells and perform flow cytometric analysis using antibodies directed against intracellular and surface hematopoietic markers to determine their phenotype. Also included is the Wright-Giemsa staining of bone marrow and spleen cells, which was done to show blast number or structural differences following drug treatment.
This protocol is robust and highly reproducible. The data shown here will aid investigators in testing the efficacy of novel therapeutic drugs. The immunocompromised RAG deleted mouse model used here is an established model for these kinds of studies, where tumor burden and survival rates can be monitored during the treatment regimen. Additionally, this method can be used to provide information on the proliferation and survival of leukemic cells and other hematopoietic cell populations throughout diagnosis.
Protocol
All methods described here have been approved by the Institutional Animal Care and Use Committee (IACUC) of the Pennsylvania State University College of Medicine.
1. Selection of animals for the study
- Maintain 18 healthy 10−12-week old male NOD.Cg-Rag1tm1Mom Il2rgtm1Wjl/SzJ (NRG) mice in a barrier environment under pathogen-free conditions.
NOTE: The common source for the right mouse strain for the study is peer-reviewed publications. The recommended age for mice is between 6 and 12 weeks-old, depending on the study. If there is no gender preference for the experiment, it is best to include both genders to avoid any gender-specific variations. Considerations about number and biological variables are explained in the discussion section.
2. Transplantation of leukemia cells
- Count the cultured luciferase labeled leukemia cells and calculate the total cell number needed based on the desired cell number/mouse (10,000 U937-Luc-tdTomato cells per mouse is needed to achieve sufficient engraftment).
NOTE: Cultured cells should not be overgrown. The density should be between 5 x 104–1 x 106 cells/mL. - Harvest the calculated cells in cell growth medium (RPMI-1640 with 10% fetal bovine serum [FBS]) in a 50 mL tube and centrifuge at 360 x g for 5 min at 4 °C.
- Dump off the supernatant and resuspend cells in Hank’s balanced salt solution (HBSS) (150 µL per mouse). If 10,000 cells per mouse are desired and there are 10 mice total, then resuspend 100,000 cells in 1,500 µL (1.5 mL) of HBSS. Place the cellular suspension on ice.
NOTE: When counting the cells, consider only viable cells, total cells. Also, prepare cells for some extra mice. For example, if 10 mice are needed for injection, prepare cells for 12−15 mice. - Fill 28 G insulin syringes with 150 µL of cell suspension. Place the mouse in a restrainer. Rub the tail gently with a 70% alcohol pad to clean the injection site.
NOTE: The mouse restrainer should be thoroughly cleaned with disinfectant (Table of Materials). - Hold the distal tail to dilate the vein. Inject the cells into the tail vein once the vein is dilated enough.
NOTE: Heat lamps can be used to dilate the tail veins. Cells settle down at the bottom of the tube, so mix cells before each injection for consistency. - After the injection, dispose of the syringe and needle into a sharps container. Press the tail to secure clotting and prevent bleeding. Lead the mouse into the cage and monitor for a few seconds to make sure there is no bleeding.
3. Imaging, randomization, and initialization of the treatment
- Depending on the cell line and animal model, perform the first animal imaging 3−12 days post-engraftment.
- Transfer the animals to the imaging room following the IACUC protocol. Record the whole bodyweight of each animal before the anesthesia.
- Turn on the anesthesia system and charge the anesthesia chamber. Set an anesthesia wheel following the manufacturer’s instructions for healthy mice (2.5%). However, set it lower (i.e., 1−1.5%) for sicker mice to deliver less anesthetic.
- Take the mice and inject them with 5 µL/g of 30 mg/mL stock luciferin intraperitoneally with a 28 G insulin syringe. Determine the amount of luciferin per mouse from the bodyweights recorded on the sheet (e.g., 24 g mouse = 120 µL, 27 g mouse = 130 µL).
NOTE: Always round the drug volume down (e.g., if the study has a 27 g mouse and a 26 g mouse, they both will be dosed at 130 µL). - As the mice are injected, move them into the anesthesia induction chamber. Once they are immobile, move them one by one into the imaging cabinet to deliver a maintenance dose of isoflurane to the mice through nosecones in the imager.
NOTE: Induction in a chamber generally requires 5% isoflurane, and maintenance (chamber or nose cone) requires 1−2% (flow rate = 0.5−1 L/min). - Position mice on a ventral position for imaging and take the images within 7−15 min of the luciferin delivery.
NOTE: Optimization of imaging time is recommended if the model is being tested for the first time. Do not open the door of the imager while an image is being taken. This might cause major issues with the machine, and the whole program may need to be restarted and reinitialized. - After acquiring the images, remove the mice from the imaging chamber and place them back into their home cage. Lay them gently on the top rack of the cage to recover from the anesthesia. Lid the cage while they recover so that they do not stumble off the cage and fall to the floor, resulting in injury or death. Once recovered and awake, place the mice gently back inside of their home cage.
- Analyze the whole-body bioluminescence images of the mice using the imaging software (Table of Materials) to free draw around the entire mouse body, excluding the tail. Region of interest (ROI) values will be automatically provided after drawing free lines surrounding each mouse. Rank the mice depending on their ROI signal levels. Evenly disperse mice among all groups based on their ROI to yield as even a group average as is possible for the entire study.
- Exclude the mice with signals that are either too high or too low. Mark (e.g., ear punch) each mouse in order to follow up the individual mouse in each study group.
- Start the treatment by following the study’s planned treatment regimen (e.g., see Figure 1C).
NOTE: Care must be taken to follow manufacturer’s instructions for the solubility, storage, and handling of each treatment agent. For example, daunorubicin (DNR) was reconstituted in saline and aliquots were stored at 4 °C. Cytarabine (Ara-C) was reconstituted in saline and stored at 25 °C. DNR (1 mg/kg) was delivered intravenously (IV) and Ara-C (25 mg/kg) was delivered intraperitoneally (IP). Store the drugs in daily-based volumes to avoid freeze-thaw cycles and maintain their stability. - Image the mice throughout the study to evaluate the treatment response following steps 3.2−3.8.
4. Blood collection (tail vein puncture)
NOTE: Blood collection should be done once per week during the study to monitor abnormal blast cells as well as complete blood count (CBC) levels for toxicity.
- Place the mouse tail first into the restrainer and place the restraint plug in the tube, flush with the mouse’s nose so that the mouse becomes immobile without being crushed. Tighten the plug so that the mouse cannot move.
NOTE: It may be useful to dip the tail in warm water to increase vasodilation prior to starting. - Wipe the tail with an alcohol pad to remove any debris. Locate one of the lateral tail veins.
- Insert a 25 G needle attached to a 1 mL syringe into the lateral tail vein, approximately 1/3 of the way down the tail from the base of the tail. Insert the needle at a ~5° angle, almost parallel to the tail, with the beveled end pointed up.
- Remove the needle and place the capillary tube up against the blood as it wells up. Depending on the blood flow, it could take several seconds for the blood to travel down the capillary into the tube. Once the blood begins to drop into the tube, periodically flick the tube gently to mix with EDTA and prevent the blood from clotting.
- Collect at least 50 µL of blood.
NOTE: Collecting less blood will make it difficult to perform the CBC counts. If blood flow is too slow, try puncturing the tail vein with the needle higher up towards the base of the tail. It is also effective to try the other lateral tail vein. Do not puncture the tail more than 3x in one sitting to prevent undue stress. - Once the blood is collected, take the capillary top off and close the lid of the tube. Gently flick the tube for 30 s to further mix and prevent clotting. If the blood clots, it cannot be used for CBC counts. Apply pressure to the tail at the puncture location for 30 s to stop the bleeding.
- Unscrew the plug and remove the mouse from the restrainer. Monitor the mouse to make sure there is no further bleeding from the tail. If bleeding continues, apply styptic powder to the puncture site.
NOTE: Clean the restrainer when changing mice to limit pheromone-induced stress and contamination. If desired, submandibular or saphenous vein bleeding can also be used10.
5. Euthanasia
NOTE: Follow the institution-approved standard procedure for mice euthanasia.
- When mice exhibit distinct signs of being moribund (i.e., hind limb paralysis, lethargy, inability to right itself, more than 10% weight loss, hunching, immobility), perform euthanasia.
NOTE: This will occur typically around day 22 postengraftment for the U937-Luc-dsRed mouse model. - For euthanasia by CO2, place the mice in the cage and place the lid attached to the CO2 regulator onto the top of the cage. Ensure that the cage is sitting flush with the top of the cage, and the air vent holes are not blocked. Open the valve on the CO2 tank to start the flow. Adjust the CO2 regulator to 1−2 L/min.
NOTE: Allow the empty chamber to charge for 2 min before placing the mice in. - Once the mice have become unconscious, as indicated by lack of movement and rapid breathing, increase the CO2 regulator knob to 4−5 L/min. Continue to monitor the mice until 1−2 min after the last visual breath. Turn the CO2 regulator valve to 0 and close the CO2 tank valve. Take the mice out of the chamber.
- Perform a secondary form of euthanasia (e.g., cervical dislocation, cardiac puncture, or removing major organs) to each mouse to ensure death.
- To perform cervical dislocation, place two fingers on either side of the mouse’s neck. While applying pressure to the neck, pull the tail up at a 45° angle until a separation of the joints in the neck is felt.
6. Leukemia cell isolation from organs (bone marrow and spleen)
- Prepare the work area by spraying the cork or lid of a polystyrene foam box surface with ethanol, covering with paper towels, and spraying with more 70% ethanol. Place a bench pad underneath the surface to limit depositing debris on the surrounding areas.
- If blood is needed for CBC and serum analysis, perform a cardiac puncture immediately after CO2 euthanasia.
- Place the mouse on its back. Place the 25 G needle attached to a syringe bevel up between the ribs on the left side of the mouse, at the level of the elbow, approximately 5 mm into the chest cavity, to puncture the heart.
- Gently pull back the plunger. If the needle is in the heart, blood will flow into the syringe. If there is no blood, pull the needle out slowly until blood starts to flow into the syringe. Keep the needle still once the blood starts to flow into the syringe.
NOTE: If unable to collect blood after several attempts, open the body cavity as described below to access the chest cavity to collect blood. - Once the blood has filled the syringe, pull the plunger back again slowly. The heart will need time to fill back up, and it may take a few seconds to get the blood flowing again. Place approximately 50 µL of blood into an EDTA tube and gently flick the tube to prevent clotting.
NOTE: The rest of the blood can be placed in a microcentrifuge tube for serum analysis. The blood for serum analysis can be left to clot.
- Prepare 6 well plates by adding 5 mL of cold phosphate-buffered saline (PBS) to each well with a 0.45 µm cell strainer. Place the 6 well plates on ice.
- Place the mouse on its back and spray the ventral surface with 70% ethanol until the fur is wet. Secure the mouse to the surface by placing a 20 G needle through the tail, ~1 cm from the base of the tail, and one needle through the neck of the mouse, just below the jaw, to allow access to the internal organs, as well as the leg bones, for bone marrow collection.
- Using forceps, pull the fur up and make a 1 cm cut through the first layer of skin using scissors. There will be another layer of fascia visible just underneath the cut.
- Pull up on the second layer of fascia and make an incision to expose the internal organs. While pulling up on the skin, make a midline cut through both layers of fascia towards the head, ending at the bottom of the sternum. Make a similar cut towards the tail of the mouse, ending in line with the hips.
- At the end of the caudal cut, make a 1 cm cut down the inside of the thigh on each leg. At this point, it may be helpful to pin back any excess skin to better visualize the internal organs. The spleen is located on the mouse’s left flank, underneath the stomach, next to the kidney. To remove the spleen, gently lift it using the forceps and cut away any connective tissue.
- Place the spleen in the cell strainer and using the back of a 1 mL syringe plunger, mash the spleen into the strainer until it becomes a single cell suspension. Mash the spleen until there is no red tissue left in the strainer, only white tissue.
- While the spleen suspension is on ice, start to remove the bone marrow by taking the previous incision on the thigh, from step 6.7, down to the ankle of the mouse. Continue that incision around the ankle and pull the skin away from the leg muscle to expose the upper thigh.
- Cut the muscle from the back of the leg bones starting from the ankle up to the pelvis. Find the head of the femur and feel for the separation between the pelvis and the head of the femur. Cut through the separation to detach the leg from the body.
NOTE: If more cells are needed for analysis, the forelimbs, spine, and iliac crest may also be harvested. - Remove the tissue from the tibia by twisting the ankle and gently pull up to remove the ankle and tissue from the tibia. For the remaining tissue, use a gauze pad or paper towel to gently pull away from the tissue without breaking the bones. Place the tibia and femur into the 6 well plate and repeat on the other leg.
- Cut the tips of the bones off and take up a few milliliters of PBS in the dish into a syringe attached to a 25 G needle. Guide the needle tip into the opening of the bone and flush out the bone marrow. Continue to flush the bones until water flushes through the bone cavity without resistance. Small pieces of bone marrow or flakes will be visible in the collection tube. Transfer the spleen and bone marrow contents to respective 15 mL conical tubes and fill to 15 mL with cold PBS.
NOTE: With advanced engraftment, there will be very little red in the bone marrow. In this case, thoroughly scrape the inside of the bones to remove as many cells as possible.- Alternatively, crush the bones with a mortar and pestle to remove the cells. Before transferring the bones to the 15 mL conical tube, thoroughly resuspend the cells, and pipette through nylon mesh to remove any bone particles. Rinse the bones with 2 mL of cold PBS to remove any cells left in the mortar, and pipette through the strainer. Fill the conical tube to 15 mL with cold PBS.
- Centrifuge at 360 x g for 5 min. Aspirate the supernatant and resuspend the pellet thoroughly in 3 mL of 1x red blood cell lysis buffer (Table of Materials).
- Incubate the suspension for 8 min at room temperature (RT), then dilute by filling the tube with cold PBS up to 15 mL. Centrifuge again at 360 x g for 5 min. The pellet should be white, with no red blood cells left.
- Aspirate the supernatant and resuspend the pellet in 1 mL of cold PBS or fluorescence-activated cell sorting (FACS) buffer (Table of Materials).
7. Cell surface and intracellular immunofluorescence staining of isolated animal cells
- Cell surface antigen staining
- Count out 0.5−1 x 106 cells per sample collected in step 6.15 and suspend them in 3 mL of FACS buffer in a 5 mL 12 x 75 mm FACS tube. Wash the cells 2x in 3 mL of FACS buffer and centrifuge at 360 x g for 5 min at 4 °C.
- Discard the supernatant (i.e., with a vacuum pump/Pasteur pipette or simply by washing off the pellet) and resuspend the pellet in its residual volume by gently tapping on the benchtop. Then add 1 µL of human Fc block (mouse anti-human Fc receptor antibody, 1:100) and 1 µL of mouse Fc block (rat anti-mouse CD16/32 antibody, 1:100) per sample to block nonspecific Fc-gamma receptor (FcgammaR)-mediated binding of antibodies. Mix the tubes and incubate them on ice for a minimum of 15 min.
NOTE: Always keep tubes on ice while working to prevent low viability. No protection from light is needed up to this step. - Prepare an antibody cocktail containing anti-mouse CD45, anti-human CD45, anti-human CD33, viability dye (i.e., 7-aminoactinomycin D [7-AAD] or fixable viability dye [FVD]), and fluorescence of cells (i.e., tdTomato or yellow fluorescent protein [YFP]) for the cell surface staining in the FACS buffer (Table 1).
NOTE: Use when two or more different staining reagents conjugated with brilliant fluorescent polymer dyes (e.g., BV421 anti-human CD33 and BV605 anti-mouse CD45) need to be used. It provides better data interpretation and helps to prevent any fluorescent dye interactions that may cause staining artifacts. An example panel for U937-tdTomato animal studies is shown in Table 1. When making the flow cytometry antibody panel, it is important to consider the fluorescence of the cells to be tested, if any. Viability dye plays a crucial role in excluding dead cells to avoid any false positive or negative results. - Add the antibody cocktail and incubate the samples on ice or at 4 °C for 30 min in the dark (i.e., cover with aluminum foil).
NOTE: The protection of samples from light becomes very important after this step. Working in a dark environment is ideal. If that is not possible, minimize the light exposure by covering the tubes with aluminum foil in between steps, for example. - Add 0.5−1 mL of FACS buffer and centrifuge the tubes at 300 x g at 4 °C for 5 min. Dump the supernatant and repeat this step.
NOTE: If intracellular staining is desired, use PBS (azide and serum/protein-free) instead of FACS buffer.- Calculate the engraftment percentage by obtaining the ratio of human CD45 (white blood cells) positive cells to whole CD45 positive cells (human CD45+ and mouse CD45+ cells) using the following formula:
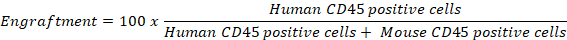
- Calculate the engraftment percentage by obtaining the ratio of human CD45 (white blood cells) positive cells to whole CD45 positive cells (human CD45+ and mouse CD45+ cells) using the following formula:
- Resuspend the stained cells in 300 μL of cold FACS buffer (for 7-AAD) or PBS (for FVD) and 0.1−0.5 μL/mL of viability dye (FVD or 7-AAD). Incubate for 5−10 min on ice or at RT while protected from the light.
NOTE: If intracellular staining is desired, use FVD instead of 7-AAD in PBS (azide and serum/protein-free). - Continue with the intracellular immunofluorescence staining protocol if desired. Otherwise, perform flow cytometry in 1 h.
- Intracellular immunofluorescence staining
NOTE: First, perform cell surface antigen staining as described in section 7.1, then follow the steps below to stain intracellular antigens.- Fixation of the cells
- Warm the fixation buffer (Table of Materials) in a water bath. Aliquot 0.5 mL of buffer per 1 x 106 cells and warm to 37 °C. After two washes in PBS (azide and serum/protein-free) as in step 7.1.5, dump off the supernatant and resuspend the pellet in 0.5 mL/tube prewarmed fixation buffer for 20 min at RT in the dark.
- Centrifuge at 350 x g for 5 min at 4 °C or 25 °C and discard the supernatant.
NOTE: The cells are now fixed. If short- or long-term storage is desired, the experiment can be stopped at this point. For short-term storage, wash cells 1x with FACS buffer, resuspend in FACS buffer, and store at 4 °C. For long-term storage, resuspend in freezing medium (90% FBS and 10% dimethyl sulfoxide [DMSO]) at -80 °C. Otherwise, continue with the permeabilization protocol.
- Permeabilization of cells
- Dilute 10x intracellular staining permeabilization wash buffer (Table of Materials) to 1x in distilled water.
- Resuspend the fixed cells in 1 mL of intracellular staining permeabilization wash buffer and centrifuge at 350 x g for 5 min. Repeat this step. After the second wash, make “Unstained” or “Isotype” control tubes. Label control tube as “Unstained” or “Isotype” and resuspend in FACS buffer. Control tube is now ready, no further staining step is needed. Put the control tube on ice.
- Intracellular antigen staining
- Resuspend fixed, permeabilized cells in residual intracellular staining permeabilization wash buffer and add the antibody of interest (i.e., PE-anti-pSTAT3-tyr705 or PE-anti-BTK) into the tubes and incubate for 20 min in the dark at RT. Wash 2x with 1 mL of intracellular staining permeabilization wash buffer and centrifuge at 350 x g for 5 min at 4 °C.
- Resuspend fixed, permeabilized, and intracellularly labeled cells in 300 µL of FACS buffer and analyze with appropriate controls.
- Fixation of the cells
8. Wright-Giemsa staining of bone marrow and spleen cells
- Proceed with the Wright-Giemsa staining protocol once the bone marrow and spleen cells are suspended (step 6.15).
- Preparation of cells for cytocentrifuge
- Prepare a cell suspension of no more than 2−5 x 105 cells in 200 μL of PBS or PBS with 2% FBS in 1.5 mL tubes. Prelabel the microscope slides (i.e., Control, U937, bone marrow [BM], NSG, Day 20, and date frozen).
- Load the slide clip with a filter card and sample chamber, followed by loading the cell suspension into the chamber.
NOTE: Make sure the chamber and filter card are aligned. Add the cell suspension slide, chamber, and slide clip set-up has been placed in the cytocentrifuge. - Cytocentrifuge the chambers at 600 x g for 5 min.
- Remove the slide from the chamber carefully by holding the slide clip firmly and pushing the spring against the slide to release it from the hooks. Then move the spring away from the hooks and allow the sample chamber to be released from the clip. Air-dry the slides and proceed with the Wright-Giemsa staining protocol.
- Wright-Giemsa staining of slides
- Stain slides with Wright-Giemsa staining solution (i.e., dip in staining solution or flood slides with staining solution on a staining rack) for 1 min.
NOTE: If staining on a rack is preferred, add sufficient Wright-Giemsa solution to cover the entire surface. - Dip the slides in distilled water for 2 min (or longer for darker staining). Rinse the slides with distilled water, and air-dry.
- Mount the slides with a coverslip using the mounting medium and examine the slides under the microscope.
- Stain slides with Wright-Giemsa staining solution (i.e., dip in staining solution or flood slides with staining solution on a staining rack) for 1 min.
Results
We developed a model to study the standard of care chemotherapeutic regimen for AML in a mouse model. Luciferase and tdTomato-expressing U937 cells were cultured to allow a few passages. The luciferase activity of the cells was checked using the BLI system and found to be highly active (Figure 1A). Cells were observed under a fluorescence microscope to confirm the tdTomato expression (Figure 1B). NRG mice were injected intravenously with U937-Luc-tdTomato cells,...
Discussion
Critical steps in the protocol
Characterization of the mouse model: Each cell line and primary cell-derived murine xenograft has a unique disease and host-specific factors that influence the time and characteristics of engraftment. Time to engraftment is usually defined as time taken for the bone marrow to have 25% blast cells or have bioluminescence signaling at least two logs higher than background or non-tumor bearing mice. It is critical to characterize the mouse model prior to starting an expe...
Disclosures
The authors declare that they have no competing interests. Mark Kester is the Chief Medical Officer of Keystone Nano, Inc. Thomas P. Loughran, Jr. is on the Scientific Advisory Board and has stock options for both Keystone Nano and Bioniz Therapeutics. There are no conflicts of interest with the work presented in this manuscript.
Acknowledgements
This study was funded by the Kenneth F Noel Memorial Fund (D.F.C.), Delbert J. McQuaide Cancer Research Fund (A.S.), Austin R. Orwan Memorial Research Fund (A.S.), The Penn State Cancer Institute (PSCI) and the National Institutes of Health (NIH) under the National Cancer Institute (T.P.L.) (P01CA171983). This work was supported by Penn State Clinical and Translational Sciences KL2 award (KL2 TR002015) to CG. The authors thank the staff of the Penn State Cancer Institute, Department of Comparative Medicine, Bioluminescence Imaging, Flow Cytometry, and Four Diamonds Developmental Therapeutic Preclinical Core facilities at Penn State University College of Medicine.
Materials
| Name | Company | Catalog Number | Comments |
| 1 mL Syringe | Fisher Scientific | 309659 | |
| 1.5 inch short bevel 20 g needle | Fisher Scientific | 305179 | |
| 1.5 mL microcentrifuge tube | Fisher Scientific | 02-682-002 | |
| 13 mm Single Ring slide | Fisherbrand | 22-037-241 | |
| 15 mL Polypropylene Conical Tube | FALCON | 352097 | |
| 1X RBC Lysis Buffer | Fisher Scientific | 501129751 | |
| 2 mL microcentrifgue tube | Fisher Scientific | 05-408-138 | |
| 4.5 inch blunt/straight tip scissors | Fisher Scientific | 28251 | |
| 4.5 inch serated straight forceps | VWR | 82027-440 | |
| 5 mL, 12x75 mm round bottom test tubes (flow tubes) | Corning | 352008 | |
| 5/8 inch sterile 25 g needle | Fisher Scientific | 305122 | |
| 6-well Non Treated Cell Culture Plate | USA Scientific | CC7672-7506 | |
| 7AAD | Biolegend | 420404 | |
| Alcohol Prep | COVIDIEN | 6818 | |
| Aluminum Foil | VWR | 89107-726 | |
| AutoFlow IR Water-Jacketed CO2 Incubator | NUAIRE | Model no. NU-8700 | |
| Blood Collection | RAM Scientific | 76011 | |
| Brilliant Stain Buffer | BD Biosciences | 563794 | |
| BV421-human CD33 | Biolegend | 366622 | |
| BV-650 mouse CD45 | BD Biosciences | 563410 | |
| Cell Analyzer | EMD Millipore Corparation | N/A | |
| Cell Strainer | FALCON | 352350 | |
| Centrifuge machine | BECKMAN COULTER | 605168-AC | |
| Count & Viability Kit | EMD Millipore Corparation | MCH100102 | |
| Cytocentrifuge | Fisher Scientific | A78300003 | |
| Cytoclip Slide Clip | Fisher Scientific | 59-910-052 | |
| Dimethyl Sulfoxide (DMSO) | EMD Millipore Corparation | 67-68-5 | |
| Disposable Centrifuge Tube | Fisher Scientific | 05-539-8 | |
| D-Luciferin-Sodium Salt | GoldBio | LUCNA-1G | |
| FACS buffer (PBS with 2% FBS (Heat-inactivated)) | N/A | N/A | |
| Filter Cards for Cytospin | Fisher Scientific | 22-030410 | |
| FITC-human CD45 | Biolegend | 304014 | |
| Fixable Viability Dye | Thermo Fischer | 65-0864-14 | |
| Fixation Buffer | Biolegend | 420801 | |
| Flow cytometer | BD Biosciences | N/A | |
| Flow Cytometry analysis software | FlowJo, LLC | Version 10 | |
| Freezing Medium (90% FBS + 10% DMSO) | N/A | N/A | |
| Graduated Tips | USASCIENTIFIC | 10/20 µL (1110-3700), 200 µL (1111-1700), and 1000 µL (1111-2720) | |
| Hank’s Balanced Salt Solution (HBSS) | Gibco | 14025092 | |
| Heat Inactivated Fetal Bovine Serum | Atlanta Biologicals | H17112 | |
| In Vivo Imaging system | Perkin Elmer | CLS136331 | |
| Insulin Syringes | BD | 329461 | |
| Inverted Microscope | Olympus | CKX31 | |
| Isoflurane | VEDCO | NDC 50989-150-15 | |
| LABGARD CLASS II TYPE A2 BIOLOGICAL SAFETY CABINET | NUAIRE | Model no. NU-425-400 | |
| Living Image Software – IVIS Lumina Series | Perkin Elmer | 128110 | |
| Low Flow CO2 Regulator | E-Z Systems | EP-1305 | |
| MB-10 tablets, sterilant | Quip Laboratories | MBTAB75BX | |
| Micro cover glass | VWR | 48366 205 | |
| Mounting medium | Fisher Scientific | SP15-100 | |
| Mouse anti-human Fc receptor antibody | BD Biosciences | 564220 | |
| Mouse cage lid for euthanasia | E-Z Systems | E-20028 | |
| NOD.Cg-Rag1tm1Mom Il2rgtm1Wjl/SzJ (NRG) | Jackson laboratory | 7799 | |
| ntracellular Staining Permeabilization Wash Buffer (10X) | Biolegend | 421002 | |
| PE-anti-BTK | Biolegend | 558528 | |
| PE-anti-pSTAT3-tyr705 | Biolegend | 651004 | |
| Penicilllin Streptomycin Solution, 100X | CORING | 30-002-CI | |
| Phosphate-Buffered Saline (PBS) | CORNIG cellgro | 21-040-CV | |
| Pipet controller | DRUMMOND Scientific | 109883 | |
| Pipette | Eppendord Research | 2.5 (O24694B), 10 (O31418B), 20 (O24694B), 100 (O337778), 200 (O26279B) and 1000 (O40665B) | |
| Rat anti-mouse CD16/32 antibody | BD Biosciences | 553142 | |
| RBC Lysis Buffer (10X) | Biolegend | 420301 | |
| Refrigerated Centrifuge | NuAire | NU-C200R | |
| Reusable sample chamber | Fisher Scientific | 5991040 | |
| RPMI-1640 medium | CORING | 10-040-CV | |
| Serological pipet | FALCON | 5mL (357543), 10mL (357551), 25mL (357535) | |
| Styptic Powder | Fisher Scientific | NC1577028 | |
| Tailveiner for mouse | Agnthos | TV-150 | |
| Trypan Blue Solution | Corning | 25900051 | |
| Wipes | Fisher Scientific | 34155 | |
| Wright-Giemsa Stain | VWR | 10143-106 |
References
- Lowenberg, B., Rowe, J. M. Introduction to the review series on advances in acute myeloid leukemia (AML). Blood. 127 (1), 1 (2016).
- De Kouchkovsky, I., Abdul-Hay, M. Acute myeloid leukemia: a comprehensive review and 2016 update. Blood Cancer Journal. 6 (7), 441 (2016).
- . Cancer Stat Facts: Leukemia - Acute Myeloid Leukemia (AML) Available from: https://seer.cancer.gov/statfacts/html/amyl.html (2019)
- Yang, X., Wang, J. Precision therapy for acute myeloid leukemia. Journal of Hematology & Oncology. 11 (1), 3 (2018).
- Zhang, Z., Tang, W. Drug metabolism in drug discovery and development. Acta Pharmaceutica Sinica B. 8 (5), 721-732 (2018).
- Sharpless, N. E., Depinho, R. A. The mighty mouse: genetically engineered mouse models in cancer drug development. Nature Reviews: Drug Discovery. 5 (9), 741-754 (2006).
- Denayer, T., Stöhr, T., Van Roy, M. Animal models in translational medicine: Validation and prediction. New Horizons in Translational Medicine. 2 (1), 5-11 (2014).
- Kuzu, O. F., Nguyen, F. D., Noory, M. A., Sharma, A. Current state of animal (mouse) modeling in melanoma research. Cancer Growth and Metastasis. 8, 81-94 (2015).
- Yada, E., Wada, S., Yoshida, S., Sasada, T. Use of patient-derived xenograft mouse models in cancer research and treatment. Future Science OA. 4 (3), 271 (2017).
- Cheung, S. T., Shakibakho, S., So, E. Y., Mui, A. L. Transfecting RAW264. 7 cells with a luciferase reporter gene. Journal of Visualized Experiments. (100), e52807 (2015).
- Li, J., Low-Nam, S., Garg, S. Generation of Luciferase-expressing Tumor Cell Lines. Bio-protocol. 8 (8), 2817 (2018).
- Barve, A., et al. Comparative utility of NRG and NRGS mice for the study of normal hematopoiesis, leukemogenesis, and therapeutic response. Experimental Hematology. 67, 18-31 (2018).
- Lee, E. M., et al. Efficacy of an Fc-modified anti-CD123 antibody (CSL362) combined with chemotherapy in xenograft models of acute myelogenous leukemia in immunodeficient mice. Haematologica. 100 (7), 914-926 (2015).
- Verhagen, H. J., et al. IGFBP7 Induces Differentiation and Loss of Survival of Human Acute Myeloid Leukemia Stem Cells without Affecting Normal Hematopoiesis. Cell Reports. 25 (11), 3021-3035 (2018).
- Xie, C., et al. Panobinostat enhances cytarabine and daunorubicin sensitivities in AML cells through suppressing the expression of BRCA1, CHK1, and Rad51. PLoS One. 8 (11), 79106 (2013).
- Hengst, J. A., et al. SKI-178: A multitargeted inhibitor of sphingosine kinase and microtubule dynamics demonstrating therapeutic efficacy in acute myeloid leukemia models. Cancer Translational Medicine. 3 (4), 109 (2017).
- Song, C., et al. Targeting casein kinase II restores Ikaros tumor suppressor activity and demonstrates therapeutic efficacy in high-risk leukemia. Blood. 126 (15), 1813-1822 (2015).
- Wunderlich, M., et al. Improved chemotherapy modeling with RAG-based immune deficient mice. PLoS One. 14 (11), 0225532 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved