A subscription to JoVE is required to view this content. Sign in or start your free trial.
Therapeutic Effects of Huazhuojiedu Decoction on Precancerous Lesions of Gastric Cancer by Regulating Mitophagy
In This Article
Summary
Here, we introduce a protocol that demonstrates the therapeutic efficacy of Huazhuojiedu decoction (HZJD) in alleviating precancerous lesions of gastric cancer using mitophagy regulation.
Abstract
This research aims to explore the therapeutic effect and potential mechanisms of Huazhuojiedu decoction (HZJD) for alleviating precancerous lesions of gastric cancer (PLGC) both in vivo and in vitro. HZJD is a traditional Chinese herbal formula consisting of 11 herbs. Sprague-Dawley (SD) rats were randomly divided into four subgroups: control group, model group, positive drug group, and HZJD group. Hematoxylin-eosin (H&E) staining, high iron diamine-alcian blue (HID-AB) staining, alcian blue-periodic acid Schiff (AB-PAS) staining, immunohistochemistry, immunofluorescence, RT-qPCR, and Western blot assays were performed after 10 weeks of HZJD treatment. In vitro, the cell counting kit-8 (CCK-8) and 5-ethynyl-2'-deoxyuridine (EdU) assays were used to detect cell proliferation. RT-qPCR and Western blot assays were performed to evaluate mitophagy levels. The results indicated that HZJD could retard the pathological progression in PLGC rats and reduce PLGC cell proliferation. Treatment with HZJD significantly increased the mRNA and protein expression levels of Sirt3, Foxo3a, Parkin, and LC3 II/I, while decreasing the mRNA and protein expression levels of p62 and Tomm20. HZJD was found to have the ability to reverse the decline in mitophagy activity both in vivo and in vitro. In conclusion, the study assessed the impact of HZJD and provided evidence regarding its potential molecular mechanism.
Introduction
Gastric cancer (GC) remains one of the most common malignant diseases affecting the digestive system worldwide. It is estimated that GC accounts for approximately 6% of all cancers worldwide, ranking 5th among the most frequently diagnosed cancers and 3rd among cancer-related deaths1. GC is widely recognized as a progressive, multi-step biological process. Prior to the onset of GC, the gastric mucosa often undergoes several years of precancerous lesions, referred to as precancerous lesions of gastric cancer (PLGC) stages. The Correa cascade theory is widely accepted and provides an explanation for the sequential progression from normal mucosa to chronic non-atrophic gastritis, atrophic gastritis, intestinal metaplasia (IM), dysplasia (Dys), and eventually carcinoma2. PLGC represents a crucial stage in GC development, and timely intervention and monitoring of PLGC are vital for early GC prevention.
Recent clinical evaluations and experimental research have confirmed that traditional Chinese medicine (TCM) is emerging as one of the most effective therapies for treating PLGC3,4. HZJD decoction is a TCM formula developed based on clinical experience and rooted in the TCM theory of heat and dampness removal. Previous studies have demonstrated the beneficial effects of HZJD decoction in the treatment of PLGC, particularly in relieving clinical symptoms and pathological manifestations5,6. Through network pharmacology-related studies, we identified the active components of HZJD decoction as well as their potential targets for PLGC7. The findings from a previous study indicated that HZJD decoction had the ability to enhance diversity, optimize the community structure, and increase the relative abundance of intestinal flora in PLGC rats8. Moreover, it was verified that HZJD decoction could improve PLGC by regulating intestinal microbiota and its metabolites9. In recent work, we demonstrated that HZJD decoction could regulate the dynamic balance of cell proliferation and apoptosis in PLGC cells by downregulating the expression of lnc 51736810.
An increasing number of studies have confirmed that mitophagy plays an important role in various cancers. Mitophagy selectively eliminates damaged and depolarized mitochondrion and further prevents excessive accumulation of cytotoxic reactive oxygen species (ROS) from the dysfunctional mitochondrion, which in turn inhibits tumorigenesis11. Mitophagy, the selective degradation of damaged or dysfunctional mitochondria, is intricately linked to mitochondrial oxidative phosphorylation (OXPHOS). The impairment or absence of mitophagy can lead to a shift in cellular energy metabolism towards aerobic glycolysis, a phenomenon commonly known as the Warburg effect. The increased production of lactate and ketone bodies, resulting from the Warburg effect, contributes to the construction of a tumor microenvironment that is conducive to cell proliferation12.
This study presents a comprehensive protocol for utilizing HZJD decoction as a therapeutic approach to mitigate the progression of PLGC. Through our evaluation, we observed a significant positive effect of HZJD decoction, particularly in its ability to regulate mitophagy. The study provides valuable insights into the potential molecular mechanisms of HZJD decoction in treating PLGC.
Protocol
All experimental procedures and animal care were approved by the Hebei University of Traditional Chinese Medicine Institutional Animal Care and Use Committee Guidelines (approval number: DWLL2019031) and were performed in accordance with the ethical guidelines. A total of 90 specific pathogen-free male Sprague-Dawley (SD) rats (age = 6 weeks; weight = 150-180 g; see Table of Materials) were raised at constant temperature (24 °C ± 4 °C) and humidity (50%-60%) under controlled dark/light cycle of 12 h. The rats were acclimated to the new environment for 1 week before starting the experiments.
1. Preparation for animal experiment
- Rat model of PLGC
- Randomly assign the rats to a control group (n=20) and a PLGC model group (n=70). Supply the rats in control group with a standard rodent pellet diet and water ad libitum. Feed the rats in PLGC model group with an irregular diet (1 day fasting, 1 day feeding).
- House the rats in groups of five per cage. Provide the control group with 150 g/day of feed per cage. Allow the PLGC model group free access to food for 23 h (150 g/day/per cage). After 23 h, remove the remaining feed.
- Provide the rats in PLGC model group with 1-methyl-3-nitro-1-nitrosoguanidine (MNNG) solution (200 µg/mL) for free drinking10,13. Provide the rats in the PLGC model group with 200 mL of MNNG solution per cage. Place the MNNG solution in opaque drinking bottles and replace daily.
- Gavage the rats in PLGC model group with 2% sodium salicylate after 24 h of fasting (once every 2 days by gavage). For gavage, use a silicone needle (see Table of Materials) with a minimum depth of the needle equal to 6 cm. Perform gavage consistently at the same time in the morning between 8:30 AM and 9:30 AM.
- Randomly select two rats from the PLGC model group for pathological examination at weeks 12, 16, 20 and 24 in order to assess the establishment of PLGC model. Consider the model successful when both the rats are diagnosed with PLGC by pathological evaluation as described in9.
- This modeling process lasts about 24 weeks. Allow the pathological evaluation to be performed independently by two senior pathologists. Determine the pathological diagnosis of PLGC in rats through the collaborative assessment of two pathologists. Take their diagnostic opinions into consideration and use the previous guidelines as a reference during the evaluation process14,15,16,17.
- Preparation of HZJD decoction
- Prepare HZJD decoction according to the method described in7.
NOTE: The herbs in HZJD decoction were purchased and authenticated by the Hebei Hospital of Traditional Chinese Medicine.
- Prepare HZJD decoction according to the method described in7.
- Grouping and drug Intervention
- Randomly divide PLGC model group into three subgroups: model group (n=20), positive drug group (n=20), and Huazhuojiedu decoction group (HZJD, n=20, Table 1).
- Gavage the rats in positive drug group with 0.7 mg/kg/day of vitamin B12 by gavage18. Treat the rats in HZJD group with HZJD decoction at 14.81 g/kg/day8. Administer the rats in control group and model group with distilled water (10 mL/kg). All four groups receive the intragastric administration once daily for 10 weeks.
NOTE: The daily recommended dose of HZJD decoction for humans is 142 g/day10. Using the body surface area calculation method, it is determined that the recommended dosage for rats is 6.25x that of humans. Therefore, the daily dosage of HZJD decoction for rats is 14.81 g/kg. The dosage was adjusted weekly according to the weight of the rats. The maximum gavage volume did not exceed 3 mL.
- Sample collection
- Sacrifice all rats after 24 h of fasting (with free access to water) at week 35. Anesthetize all rats with isoflurane (5% induction, 2% for maintenance, 1L/min flow rate).
- Shave the abdominal area and sterilize the skin with ethanol and iodine after confirming the loss of pain stimulation. Cut the abdominal skin from the xiphoid cartilage along the midline of the abdomen with a scalpel.
- Blunt separate the subcutaneous tissue with forceps and a scalpel until exposing the stomach. Dissect the stomach along the greater curvature with a scissor and immediately rinse it with phosphate-buffered saline (PBS).
- Unfold the stomach tissues on an ice plate. Collect gastric samples (2 mm x 2 mm) from the antrum, the corpus, and the lesser curvature of the angular region. If there are any visible lesions on the stomach, collect and process those specific portions as specimens.
- Fix gastric samples in 4% paraformaldehyde for 24 h. Place the remaining gastric tissues into cryotubes. Freeze them in liquid nitrogen, and then store them at -80 °C.
- Euthanize the rats via inhalation of carbon dioxide. Place rat carcasses in sealed bags and then put bags in the animal carcass storage cabinet.
2. Pathological examination
- Trim and smooth the fixed gastric tissues with scissors. Dehydrate tissues with a gradient alcohol series, i.e., 75% ethanol for 30 min, 85% ethanol for 30 min, 95% ethanol for 30 min, and 2x in anhydrous ethanol for 30 min each. Immerse tissues in xylene/ethanol solution (1:1) for 1 h, and 2x in 100% xylene for 30 min each.
- Pour half of the molten wax into mold, and then put the permeabilized tissues in mold quickly. Place the tissues into the other half of the molten wax. Mark the embedded box and allow the wax to completely solidify.
- Use microtome to cut into slices with a thickness of 4 µm. Place the slices on the slides.
- Place the slices 2x in 100% xylene for 10 min each, then in xylene/ethanol solution (1:1) for 10 min, 2x again in anhydrous ethanol for 5 min, then in 95% ethanol for 5 min, followed by 85% ethanol for 5 min, and finally in 70% ethanol for 5 min. Rinse the slices with running water.
- Stain the slices with hematoxylin for 5 min. Differentiate by dipping in 0.5% hydrochloric acid-ethanol for 10 s. Stain the slices with eosin for 2 min.
- Rinse the slices with distilled water. Dehydrate the slices with a gradient alcohol series, i.e., 70% ethanol for 30 s, 80% ethanol for 30 s, 95% ethanol for 30 s, anhydrous ethanol for 30 s. Permeabilize the slices with 100% xylene 2x. Seal the slices with neutral balsam.
- Mix HID solution A and HID solution B at a ratio of 50:3 to prepare the HID working solution. Stain the slices with the HID working solution for 24 h.
- Rinse under running water and then stain the slices with alcian for 20 min. Counterstain the slices with the nuclear fast red solution for 10 min. Dehydrate and seal the slices as described in step 2.6.
- Stain the slices with alcian for 20 min. Incubate the slices in 1% aqueous solution of periodic acid for 5 min. Stain the slices with Schiff for 20 min.
- Incubate the slides with hematoxylin for 2 min to stain the nuclei. Add the acidic differentiation solution (provided with the HID kit) for 5 s. Apply the Scott blue solution for 3 min to color the slides blue. Dehydrate and seal the slices as described above in step 2.6.
NOTE: The staining process should be protected from light. The temperature should be kept below 22 °C when using 1% aqueous solution of periodic acid. The slices should be dehydrated and sealed as described in step 2.6. - Examine the stained slices under an optical microscope at magnification of 10x and 20x.
3. Immunohistochemistry
- Use the slices obtained in step 2.4. Put the slices in 0.01 M sodium citrate buffer and heat for 10 min to perform antigen retrieval.
- Add 3% hydrogen peroxide for 30 min to quench the endogenous peroxidase and biotin. Block the slices with goat serum for 30 min.
- Incubate the slices with the diluted primary antibodies against sirt3 (1:200), foxo3a (1:100) and parkin (1:200) overnight at 4 °C. Substitute the primary antibodies with PBS for the negative control.
- Rinse the slices 3x for 5 min per rinse with PBS. Incubate the slices with the corresponding secondary antibody at room temperature for 1 h.
- Add 20 µL of 3,3'-diaminobenzidine (DAB) for 3 min. Dehydrate and seal the slices as described above in step 2.6.
- Image all slices at 40x magnification under the optical microscope. Use Image-Pro Plus 6.0 software to perform quantitative image analysis.
4. Immunofluorescence
- Use the slices obtained in step 3.1. Repeat step 3.3.
- Incubate the slices with the diluted primary antibodies against COX IV (1:500), and LC3 (1:500) overnight at 4 °C. Rinse the slices 3x for 5 min per rinse with PBS.
- Incubate the slices with the goat anti-rabbit IgG (1:1000) and goat anti-mouse IgG (1:1000) at room temperature for 1.5 h. Rinse the slices 3x for 5 min per rinse with PBS.
- Add DAPI staining solution and incubate at room temperature for 10 min in the dark. Rinse the slices 3x for 5 min per rinse with PBS.
- Add dropwise DAPI staining solution and incubate at room temperature for 10 min in the dark. Mount the slices with anti-fade fluorescence mounting medium.
- Visualize and photograph the slices under a fluorescence microscope (see Table of Materials) at 40x magnification.
5. Western blot analysis
- Weigh 100 mg of gastric tissues precisely. Add 1 mL of RIPA buffer solution (see Table of Materials). Grind the tissues thoroughly using a homogenizer (10,000 x g, 15 s each time for 3x).
- Place the tissue homogenate on ice for 30 min. Centrifuge at 12,000 x g for 20 min at 4 °C, and then collect the supernatant.
- Quantify the protein concentration by using the bicinchoninic acid (BCA) protein concentration determination kit (see Table of Materials). Adjust the protein concentration of the obtained supernatant to be consistent within samples.
- Prepare SDS-PAGE gel consisting of a 10% separating gel and a 5% stacking gel. Pour the 10% separation gel into the glass plate to 2/3 of the total height. Add deionized water over the gel until the gel solidifies. Pour the 5% stacking gel to fill the glass plate. Insert the electrophoresis comb.
NOTE: Care should be taken to ensure that there are no bubbles in the gel. - Mix the protein supernatant with 5x loading buffer at a ratio of 1:4. Put it in boiling water for 5 min to denature. Store it at -20 °C.
- Assemble the prepared SDS-PAGE gel in the Western blotting electrophoresis system. Add fresh electrophoresis solution (see Table of Materials). Load 20 µL of sample per well onto the gel. Start the electrophoresis at 80 V for 40 min and then switch to 120 V.
- Collect the gel and create a transfer sandwich as follows: two layers of sponge pad, two layers of filter paper, gel, polyvinylidene fluoride (PVDF) membrane (see Table of Materials), two layers of filter paper, two layers of sponge pad (from the negative pole to the positive pole). Add pre-cooled transfer buffer. Perform wet transfer at 350 mA for 2 h.
NOTE: Before assembling the transfer sandwich, the PVDF membrane should be pretreated with methanol. Additionally, the sponge, filter paper, and pretreated PVDF membrane should be immersed in the transfer buffer (see Table of Materials) in advance. - Block the membrane with 5% non-fat milk in tris-buffered saline with tween 20 (TBST) for 2 h on a shaker. Incubate the membrane overnight at 4 °C with the following diluted primary antibodies (Sirt3 = 1:500, Foxo3a = 1:1000, Parkin = 1:2000, P62 = 1:1000, LC3 = 1:1000, Tomm20 = 1:2000, β-actin = 1:5000, GAPDH = 1:5000).
NOTE: GAPDH was selected as reference for the detection of Parkin due to the molecular weight of Parkin (50 kDa) being close to that of β-actin (42 kDa). - Wash the membrane 4x with TBST for 5 min each time. Incubate it with the diluted secondary antibody (1:5000) at room temperature for 1 h.
- Drop the ECL working solution on the protein side of membranes for 2 min. Acquire the images through the chemiluminescence imaging system. Use ImageJ software to measure the greyscale values.
6. Quantitative real-time PCR analysis
- Clean all experimental devices with 0.1% diethyl pyrocarbonate (DEPC), and then sterilize them at high temperature and high pressure in advance.
- Weigh 50 mg of gastric tissues. Add 1 mL of Redzol (see Table of Materials) and grind in a homogenizer. Transfer tissue homogenate into an RNase-free centrifuge tube and let it stand at room temperature for 10 min.
- Add 0.2 mL of trichloromethane and mix thoroughly. Let it stand at room temperature for 3 min. Centrifuge at 4 °C and 12,000 x g for 15 min.
- Remove the upper aqueous phase carefully and mix it with 0.5 mL of isopropanol for 10 min. Centrifuge 12,000 x g for 10 min at 4 °C to precipitate RNA. Discard the supernatant and then wash the RNA precipitate 3x with 75% ethanol.
- Centrifuge at 7,500 x g for 5 min at 4 °C. Discard the supernatant and then dry RNA precipitate in a sterile workbench for 10 min. Dissolve the precipitate in 50 µL of DEPC.
- Reverse-transcribe 1 µg RNA into cDNA. Add 1 µL of 20x RTase mix, 4 µL of 5x RT reaction buffer (see Table of Materials) and replenish DEPC to a total volume of 20 µL. Set the reaction procedure as follows: 10 min at 25 °C, 40 min at 42 °C, 10 min at 85 °C, and then hold at 4 °C.
- Prepare amplification reaction mixture containing 2 µL of cDNA, 2 µL of forward primer, 2 µL of reverse primer (Table 2), 10 µL of 2x SYBR premix (see Table of Materials), and 4 µL of DEPC. Set the reaction procedure as follows: 95 °C for 15 s, 60 °C for 10 s, and 72 °C for 30 s (40 cycles).
- Use β-actin as an endogenous reference. Obtain the Ct values. Calculate the relative expression of target genes by using the 2-ΔΔCt method.
7. Preparation for cell experiments
- Cell model of PLGC
- Culture the GES-1 cells in Roswell Park Memorial Institute (RPMI) 1640 medium combined with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin. Culture the cells at 2 x 105 cells per well in a 6-well plate containing 1.5 mL of RPMI 1640 medium without serum and antibiotics at 37 °C with 5% CO2.
- Induce the GES-1 cells with MNNG for malignant transformation. Set the modeling arrangement as described below.
- On day 0, pre-resuscitate and passage GES-1 cells to maintain them in the logarithmic growth phase. Inoculate the log-phase GES-1 cells in culture flasks and culture them overnight.
- On day 1 and day 2, replace the MNNG (10 µM/L)-containing medium. On day 3, replace the MNNG-containing medium with drug free medium.
- On day 4 and day 6, treat the cells with MNNG (5 µM/L)-containing medium for 24 h. On day 7, replace the MNNG-containing medium with drug free medium. Remove the numerous dying and shedding cells and continue the cell culture.
- On day 8 and day 10, treat the cells with the medium containing MNNG (5 µM/L) for 24 h. On day 11, construct the cell model of PLGC (MC).
NOTE: During the modeling period, the status of the cells should be observed regularly. If the cells show a poor status, it is recommended to treat them with MNNG for different time intervals, preferably with a 2-day interval. Alternatively, the cells can be treated with a low dose of MNNG (3 µM/L). The pathological diagnosis criteria of MC cells can be referenced from previous research to assess the status of the cells accurately10,19,20,21.
- Preparation of drug-containing serum
- Feed 30 SD rats (male, 220 g) with a standard rodent pellet diet and water ad libitum until all rats weigh more than 350 g. Assign the rats randomly to positive drug-containing serum group (n=10), HZJD-containing serum group (n=10), and normal rat serum group (n=10).
- Perform drug administration in rats as described in step 1.3.2. Administer the rats 2x daily at 8:00 AM and 8:00 PM. Continue the drug intervention for 7 days.
- Collect the blood samples within 1 h after the last administration on day 8 (8:00 AM). Place the samples in anticoagulant tubes at 4 °C for 1 h. Centrifuge at 3,000 x g for 15 min at 4 °C and then collect the supernatant.
- Grouping and intervention methods of cell experiments
- Divide the cell experiments as follows: GES-1 group (GES-1), MC group (MC), positive drug group (positive drug), HZJD decoction group (HZJD), Sirt3 silencing group (si-Sirt3), negative control group (si-NC) and Sirt3 silencing combined with HZJD decoction group (si-Sirt3+HZJD).
- Treat the cells in positive drug group and HZJD group with 10% vitamin B12-containing serum and 10% HZJD-containing serum, respectively. Administer other groups with 10% normal rat serum.
NOTE: Based on previous research10, 10% drug-containing serum was selected for cell treatment. - Perform Sirt3 si-RNA (see Table of Materials) transfection for the cells in si-Sirt3 group and si-Sirt3+HZJD group. Set up a negative control group receiving an empty vector, namely, si-NC.
- Dissolve 2.5 nM siRNA power in 125 µL of DEPC to prepare siRNA solution. Dilute 5 µL of siRNA solution with 250 µL of minimum essential medium and dilute 5 µL of lipofectamine 2000 with 250 µL of minimum essential medium. Mix them together evenly.
- Add the mixture to RPMI 1640 medium (1.5 mL/per well) in a 6 well plate. Incubate at 37 °C with 5% CO2. Perform Western blot analysis to detect the protein expression level of Sirt3 at 48 h after transfection.
8. CCK-8 assay
- Prepare the cells into cell suspension when cells cover 70% of the bottom of the bottle. Use the cell counter to count the number of cells in the suspension. Adjust the concentration to ensure 2,000 cells/well, and each well contains 100 µL of RPMI 1640 medium. Seed the cells in 96-well plates and maintain at 37 °C in an incubator with 5% CO2 for 24 h. Set up 6 replicate wells for each group.
NOTE: The peripheral wells of the 96-well plates that are not being used are filled with 200 µL of PBS in order to reduce evaporation during the incubation period. - After culturing the cells for 24 h, replace the culture medium with a different drug-containing culture medium. Culture the cells for 48 h with positive drug-containing culture medium and HZJD-containing culture medium.
NOTE: The culture medium is clear. Observing the bottom of the culture bottle with light, it can be seen that the cells are connected into sheets. This phenomenon can prove that the cells have adhered to the wall. When observed under a microscope, the adherent cells extend into shuttles at the bottom of the bottle and when shaking the culture medium, the cells do not move. Cells usually adhere to the wall after 24 h of culture. - Replace the drug-containing culture medium with fresh culture medium. Add 10 µL of CCK-8 solution (see Table of Materials) to each well and incubate at 37 °C for 1 h. To avoid the formation of air bubbles, add the CCK-8 solution obliquely to the culture plate wall.
- Detect the OD value of each well using a microplate reader at 450 nm wavelength.
9. EdU cell proliferation assay
- Seed the cells in 96-well plates as described in step 7.1. Replace the culture medium with different drug-containing culture medium for 48 h.
NOTE: The optimal treatment time was confirmed to be 48 h through preliminary experiments. - Remove the medium and wash the cells with PBS. Add 100 µL of culture medium per well. Prepare the 2x EdU solution (see Table of Materials, 20 µM). Add 6 µL of the EdU stock solution to 3 mL of culture medium. Add 100 µL of the 2x EdU solution (37 °C preheated) per well and then incubate for 2.5 h.
NOTE: The EdU stock solution was diluted at a ratio of 1:500 to obtain the 2x EdU solution. The actual volume of 2x EdU solution was 2.4 mL for 24 wells. - Remove the EdU-containing medium. Add 200 µL of 4% paraformaldehyde per well at room temperature for 15 min. Remove paraformaldehyde. Rinse the cells with wash solution (3% bovine serum albumin [BSA] in PBS) 3x for 5 min each.
- Remove the wash solution. Add 200 µL of permeabilization solution (0.3% Triton-X 100 in PBS) per well at room temperature for 20 min. Remove the permeabilization solution. Rinse the cells with wash solution (3% BSA in PBS) 2x for 5 min each.
- Dissolve the additive with 1.3 mL of deionized water to complete the solution. Prepare the reaction solution containing 1.72 mL of reaction buffer, 4 µL of azide 488, 80 µL of CuSO4 and 200 µL of additive solution.
- Add 50 µL of reaction solution per well. Incubate the samples at room temperature for 30 min in the dark. Rinse the cells with wash solution 3x for 5 min each.
- Remove the wash solution. Add 200 µL of 1x Hoechst 33342 solution and incubate the samples at room temperature for 10 min in the dark.
- Acquire images using the fluorescence microscope. Use Image-Pro plus 6.0 software to perform quantitative image analysis.
NOTE: The maximum excitation and emission wavelengths of Azide 488 are 495 nm and 519 nm, respectively. The maximum excitation and emission wavelengths of Hoechst 33342 are 346 nm and 460 nm, respectively.
10. Mitophagy detection in living cells
- Seed the cells at 2 x 104 cells per well in a glass bottom 24-well plate. Culture overnight cells with RPMI 1640 medium without antibiotics at 37 °C with 5% CO2.
- Prepare the mtphagy dye working solution and lyso dye working solution (see Table of Materials). Dilute the mtphagy dye solution with Hanks' HEPES buffer to a concentration of 100 nM/L. Dilute the lyso dye solution with Hanks' HEPES buffer to a concentration of 100 µM/L.
- Remove the culture medium and rinse the cells with Hanks' HEPES buffer 2x. Add 500 µL of mtphagy dye working solution (100 nM/L) and then incubate at 37 °C for 30 min.
- Rinse the cells with Hanks' HEPES buffer 2x. Add 500 µL of lyso dye working solution (100 µM/L) and then incubate at 37 °C for 30 min.
- Rinse the cells with Hanks' HEPES buffer 2x. Determine the level of mitophagy by confocal fluorescence microscopy.
NOTE: The cells were imaged without any fixation before or after staining. The imaging process was conducted as quickly as possible to minimize any potential changes or alterations in the cellular state.
11. Statistical analyses
- Use commercial software to perform statistical analyses. Compare the differences among different groups with one-way ANOVA followed by the LSD post hoc test. Present the data as mean ± standard deviation (SD). Set the statistical significance as P<0.05.
Results
MNNG induces PLGC progression in animal model and promotes morphological transformation of GES-1 cells
Upon macroscopic observation, the gastric mucosa of rats in the control group appeared uniformly bright red, smooth, and soft, with the mucosal folds arranged in a linear pattern. In contrast to the rats in control group, the gastric mucosa of the rats in model group exhibited paleness and roughness, and the mucosal plicas were flat or even disappeared (F...
Discussion
PLGC serves as a key process in the progression from chronic gastritis to GC. TCM has been proven to be a promising treatment and intervention for PLGC in recent years24,25. In recent years, TCM has emerged as a promising approach for the treatment and intervention of PLGC. This is consistent with previous research10. The results of pathological examination revealed that HZJD decoction obviously attenuated pathological injury in PLGC rats....
Disclosures
All authors declare that they have no conflict of interest.
Acknowledgements
This project was funded by the Natural Science Foundation of Hebei Province of China (H2020423207).
Materials
| Name | Company | Catalog Number | Comments |
| 1-Methyl-3-nitro-1-nitrosoguanidine | Shanghai McLean Biochemical Technology Co., Ltd., Shanghai, CHN | R030453 | |
| 3,3’-diaminobenzidine | Beijing Zhong Shan-Golden Bridge Biological Technology Co., Ltd., Beijing, CHN | ZLI0919 | |
| Alcian blue periodic acid schiff staining reagent kit | Beijing Solarbio Technology Co., Ltd., Beijing, CHN | G1285 | |
| Anti-fade fluorescence mounting medium | Beijing Solarbio Technology Co., Ltd., Beijing, CHN | S2110 | |
| Bicinchoninic acid protein concentration determination kit | Wuhan Servicebio Technology Co., Ltd., Wuhan, CHN | G2026-1000T | |
| CCK-8 reagent kit | Boster Biological Engineering co., Ltd., Wuhan, CHN | AR1160 | |
| Confocal fluorescence microscopy | Leica Instruments Co., Ltd., Weztlar, GER | TCS-SP8SR | |
COX 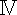 antibody antibody | Abcam Trading Co., Ltd., Shanghai, CHN | ab202554 | |
| DAPI staining solution | Beijing Solarbio Technology Co., Ltd., Beijing, CHN | C0065 | |
| DEPC | Wuhan Servicebio Technology Co., Ltd., Wuhan, CHN | G3004 | |
| EdU cell proliferation kit | Beyotime Biotechnology Co., Ltd., Shanghai, CHN | C0071S | |
| Electrophoresis solution | Boster Biological Technology co., Ltd., Wuhan, CHN | AR0139 | |
| Ethanol | Tianjin Baishi Chemical Industry Co., Ltd., Tianjin, CHN | 64-17-5 | |
| FBS | Gibco Corporation, Gaithersburg, USA | 16000044 | |
| Fluorescence microscope | Olympus Corporation, Tokyo, JPN | BH2-RFCA | |
| Foxo3a antibody | Abcam Trading Co., Ltd., Shanghai, CHN | 23683 | |
| GAPDH antibody | Wuhan Sanying Biology Technology Co., Ltd., Wuhan, CHN | 10494-1-AP | |
| Gel preparation kit | Boster Biological Technology co., Ltd., Wuhan, CHN | AR0138 | |
| GES-1 cell | Procell Life Science&Technology Co., Ltd., Wuhan, CHN | CL-0563 | |
| Goat-anti-mouse IgG | CST Biological Reagents Co., Ltd., Shanghai, CHN | 4409 | |
| Goat-anti-rabbit IgG | Abcam Trading Co., Ltd., Shanghai, CHN | ab150077 | |
| Hematoxylin-eosin staining solution | Zhuhai Beso Biotechnology Co., Ltd., Shenzhen, CHN | BA4027 | |
| High iron diamine/alcian blue staining reagent kit | Beijing Solarbio Technology Co., Ltd., Beijing, CHN | G2070 | |
| ImageJ | National Institutes of Health, Bethesda, USA | ||
| Image-Pro Plus 6.0 software | Media Cybernetics Inc., Maryland, USA | ||
| Isopropanol | Tianjin Baishi Chemical Industry Co., Ltd., Tianjin, CHN | 67-63-0 | |
| LC3 antibody | CST Biological Reagents Co., Ltd., Shanghai, CHN | 83506 | |
| Loading buffer | Boster Biological Technology co., Ltd., Wuhan, CHN | AR0198 | |
| Microplate reader | Rayto Life and Analytical Sciences Co., Ltd., Shenzhen, CHN | RT6100 | |
| Microtome | Leica Instruments Co., Ltd., Weztlar, GER | RM2245 | |
| Mitophagy kit | Dojindo Laboratories, Kyushu Island, JPN | MD01-10 | |
| Neutral balsam | Wuhan Servicebio Technology Co., Ltd., Wuhan, CHN | WG10004160 | |
| Optical microscope | Olympus Corporation, Tokyo, JPN | BH2-RFCA | |
| P62 antibody | Abcam Trading Co., Ltd., Shanghai, CHN | ab91526 | |
| Paraformaldehyde | Biosharp Life Sciences, Anhui, CHN | BL539A | |
| Parkin antibody | CST Biological Reagents Co., Ltd., Shanghai, CHN | 32833 | |
| Penicillin–streptomycin | Gibco Corporation, Gaithersburg, USA | 15140122 | |
| phosphate-buffered saline | Wuhan Servicebio Technology Co., Ltd., Wuhan, CHN | G0002-15 | |
| PVDF membrane | Millipore Corporation, Michigan, USA | IPVH00010 | |
| Redzol | SBS Genetech Co., Ltd., Beijing, CHN | FTR-50 | |
| Reverse transcription reagent kit | Igene Biotechnology Co., Ltd., Guangzhou, CHN | QP057 | |
| RIPA Buffer solution | Beijing Solarbio Technology Co., Ltd., Beijing, CHN | R002 | |
| Roswell Park Memorial Institute | Gibco Corporation, Gaithersburg, USA | 11875093 | |
| Silicone needle | Zhongke Life Biotechnology Co., Ltd., Beijing, CHN | TFEP-2 | |
| siRNA | Wuhan Genecreate Biological Engineering Co., Ltd. ,Wuhan,CHN | ||
| Sirt3 antibody | Abcam Trading Co., Ltd., Shanghai, CHN | 189860 | |
| Sodium salicylate | Shanghai Aladdin Biochemical Technology Co., Ltd., Shanghai, CHN | S104176 | |
| Sprague-Dawley rat | Beijing Huafukang Biotechnology Co., Ltd., Beijing, CHN | 110322210102553975 | |
| SYBR quantitative PCR kit | Wuhan Servicebio Technology Co., Ltd., Wuhan, CHN | G3320-15 | |
| Tomm20 antibody | Abcam Trading Co., Ltd., Shanghai, CHN | ab186735 | |
| Transferring buffer | Boster Biological Technology co., Ltd., Wuhan, CHN | AR0141 | |
| Trichloromethane | Shanghai McLean Biochemical Technology Co., Ltd., Shanghai, CHN | 821112 | |
| Tris-buffered saline with Tween 20 | Wuhan Servicebio Technology Co., Ltd., Wuhan, CHN | G0004 | |
| VDAC1 antibody | Abcam Trading Co., Ltd., Shanghai, CHN | ab15895 | |
| Xylene | Tianjin Baishi Chemical Industry Co., Ltd., Tianjin, CHN | 1330-20-7 | |
| β-actin antibody | Abcam Trading Co., Ltd., Shanghai, CHN | ab8226 |
References
- Bray, F., et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians. 68 (6), 394-424 (2018).
- He, J., et al. Helicobacter pylori infection induces stem cell-like properties in Correa cascade of gastric cancer. Cancer Letters. 542, 215764 (2022).
- Xu, W., Li, B., Xu, M., Yang, T., Hao, X. Traditional Chinese medicine for precancerous lesions of gastric cancer: A review. Biomedicine & Pharmacotherapy. 146, 112542 (2022).
- Yang, L., et al. A Systematic Review of the Mechanisms Underlying Treatment of Gastric Precancerous Lesions by Traditional Chinese Medicine. Evidence-Based Complementary and Alternative Medicine. 2020, 9154738 (2020).
- Ping, Z., Yan, W., Jing, L., Qian, J., Xin, H. The Effect of Huazhuo Jiedu recipe on Epithelial -Mesenchymal-Transition in Chronic Erosive Gastritis Patients with Zhuoduneiyun syndrome. Pharmacology and Clinics of Chinese Materia Medica. 35 (6), 154-158 (2019).
- Yan, W., Jing, L., Pan, Z. The effect of Huazhuojiedu formula on HGF/c-Met signal pathway in patients with chronic erosive gastritis. Pharmacology and Clinics of Chinese Materia Medica. 33 (02), 186-189 (2017).
- Hao, X., et al. Integrating Network Pharmacology and Experimental Validation to Investigate the Mechanisms of Huazhuojiedu Decoction to Treat Chronic Atrophic Gastritis. Evidence-Based Complementary and Alternative Medicine. 2020, 2638362 (2020).
- Zhou, P., et al. 16S rRNA sequencing-based evaluation of the protective effects of Hua-Zhuo-Jie-Du on rats with chronic atrophic gastritis. BMC Complementary Medicine and Therapies. 22 (1), 71 (2022).
- Zhou, P., et al. Determination of the protective effects of Hua-Zhuo-Jie-Du in chronic atrophic gastritis by regulating intestinal microbiota and metabolites: combination of liquid chromatograph mass spectrometer metabolic profiling and 16S rRNA gene sequencing. Chinese Medicine. 16 (1), 37 (2021).
- Hao, X., Zhou, P., Yang, Z., Yang, T., Wang, Y. The therapeutic effect of Huazhuojiedu decoction on precancerous lesions in a gastric cancer model via the regulation of lnc 517368. Journal of Ethnopharmacology. 283, 114635 (2022).
- Palikaras, K., Lionaki, E., Tavernarakis, N. Mechanisms of mitophagy in cellular homeostasis, physiology and pathology. Nature Cell Biology. 20 (9), 1013-1022 (2018).
- Zhang, C., et al. Parkin, a p53 target gene, mediates the role of p53 in glucose metabolism and the Warburg effect. Proceedings of the National Academy of Sciences of the United States of America. 108 (39), 16259-16264 (2011).
- Cai, T., et al. Protective effects of Weipixiao decoction against MNNG-induced gastric precancerous lesions in rats. Biomedicine & Pharmacotherapy. 120, 109427 (2019).
- Shah, S. C., Piazuelo, M. B., Kuipers, E. J., Li, D. AGA Clinical Practice Update on the Diagnosis and Management of Atrophic Gastritis: Expert Review. Gastroenterology. 161 (4), 1325-1332 (2021).
- Pimentel-Nunes, P., et al. Management of epithelial precancerous conditions and lesions in the stomach (MAPS II): European Society of Gastrointestinal Endoscopy (ESGE), European Helicobacter and Microbiota Study Group (EHMSG), European Society of Pathology (ESP), and Sociedade Portuguesa de Endoscopia Digestiva (SPED) guideline update 2019. Endoscopy. 51 (4), 365-388 (2019).
- Nagtegaal, I. D., et al. The 2019 WHO classification of tumours of the digestive system. Histopathology. 76 (2), 182-188 (2020).
- Kushima, R. The updated WHO classification of digestive system tumours-gastric adenocarcinoma and dysplasia. Der Pathologe. 43 (1), 8-15 (2022).
- Yang, P., et al. Weipiling decoction alleviates N-methyl-N-nitro-N'-nitrosoguanidine-induced gastric precancerous lesions via NF-κB signalling pathway inhibition. Chinese Medicine. 17 (1), 104 (2022).
- Xu, J., et al. Xiao Tan He Wei Decoction reverses MNNG-induced precancerous lesions of gastric carcinoma in vivo and vitro: Regulation of apoptosis through NF-κB pathway. Biomedicine & Pharmacotherapy. 108, 95-102 (2018).
- Li, S. Study on the mechanism of regulating NF-kB activity and inhibiting the migration of gastric "inflammation-cancer" transformed cells by invigorating spleen, removing blood stasis and detoxifying. Guangzhou University of Chinese Medicine. , 17-19 (2021).
- Liu, H. Research Progress of Cell Models of Gastric Cancer Precancerous Lesions. Traditional Chinese Drug Research and Clinical Pharmacology. 27 (4), 592-596 (2016).
- Zheng, J., et al. Chronic stress accelerates the process of gastric precancerous lesions in rats. Journal of Cancer. 12 (14), 4121-4133 (2021).
- Yu, W., et al. Sirt3 deficiency exacerbates diabetic cardiac dysfunction: Role of Foxo3A-Parkin-mediated mitophagy. Biochimica et Biophysica Acta. Molecular Basis of Disease. 1863 (8), 1973-1983 (2017).
- Cao, Y., et al. Efficacy of Banxia Xiexin decoction for chronic atrophic gastritis: A systematic review and meta-analysis. PLoS One. 15 (10), e0241202 (2020).
- Yin, J., et al. Weiqi Decoction Attenuated Chronic Atrophic Gastritis with Precancerous Lesion through Regulating Microcirculation Disturbance and HIF-1α Signaling Pathway. Evidence-Based Complementary and Alternative Medicine. 2019, 2651037 (2019).
- Zhang, J., Wang, X., Wang, F., Tang, X. Xiangsha Liujunzi Decoction improves gastrointestinal motility in functional dyspepsia with spleen deficiency syndrome by restoring mitochondrial quality control homeostasis. Phytomedicine. 105, 154374 (2022).
- Shida, M., et al. Impaired mitophagy activates mtROS/HIF-1α interplay and increases cancer aggressiveness in gastric cancer cells under hypoxia. International Journal of Oncology. 48 (4), 1379-1390 (2016).
- Zhou, X. Y., et al. Inhibition of autophagy blocks cathepsins-tBid-mitochondrial apoptotic signaling pathway via stabilization of lysosomal membrane in ischemic astrocytes. Cell Death & Disease. 8 (2), e2618 (2017).
- He, R., Peng, J., Yuan, P., Xu, F., Wei, W. Divergent roles of BECN1 in LC3 lipidation and autophagosomal function. Autophagy. 11 (5), 740-747 (2015).
- Fernández-Coto, D. L., et al. Quantitative proteomics reveals proteins involved in the progression from non-cancerous lesions to gastric cancer. Journal of Proteomics. 186, 15-27 (2018).
- Ding, D., et al. Post-translational modification of Parkin and its research progress in cancer. Cancer Communications. 39 (1), 77 (2019).
- Wang, Y., et al. The Role of Mitochondrial Dynamics and Mitophagy in Carcinogenesis, Metastasis and Therapy. Frontiers in Cell and Developmental Biology. 8, 413 (2020).
- Gehrke, S., et al. PINK1 and Parkin control localized translation of respiratory chain component mRNAs on mitochondria outer membrane. Cell Metabolism. 21 (1), 95-108 (2015).
- Fu, Z. J., et al. HIF-1α-BNIP3-mediated mitophagy in tubular cells protects against renal ischemia/reperfusion injury. Redox Biology. 36, 101671 (2020).
- Jia, Q., et al. Hesperidin promotes gastric motility in rats with functional dyspepsia by regulating Drp1-mediated ICC mitophagy. Frontiers in Pharmacology. 13, 945624 (2022).
- Chen, Y. The N-alkylamides induces MGMT gene hypomethylation in gastric epithelium cells malignant transformation. Zhejiang University. , 33-35 (2015).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved