A subscription to JoVE is required to view this content. Sign in or start your free trial.
A Live-cell Image-Based Machine Learning Strategy to Monitor Pluripotent Stem Cell Differentiation
* These authors contributed equally
In This Article
Summary
Available pluripotent stem cell (PSC)-to-functional cell differentiation systems are currently impeded by problems of severe line-to-line and batch-to-batch variability. Here, using cardiac differentiation as the main example, we present a protocol to intelligently monitor and modulate the process of PSC differentiation based on image-based machine learning.
Abstract
Pluripotent stem cell (PSC) technologies have been widely used in drug discovery, disease modeling, and regenerative medicine. However, available PSC-to-functional cell differentiation systems are impeded by problems of severe line-to-line and batch-to-batch variability. Precise control of cell differentiation in real time is therefore important. In this protocol, we describe a non-invasive and intelligent strategy that overcomes the variability in cell differentiation by using bright-field image-based machine learning. Taking PSC-to-cardiomyocyte differentiation as an example, this methodology provides detailed information for control of the initial PSC state, early assessment and intervention in differentiation conditions, and elimination of the misdifferentiated cell contamination, together realizing consistently high-quality differentiation from PSCs to functional cells. In principle, this strategy can be extended to other cell differentiation or reprogramming systems with multiple steps to support cell manufacturing, as well as to further our understanding of the mechanisms during cell fate conversion.
Introduction
Pluripotent stem cells (PSCs) possess the remarkable ability to differentiate into many types of cells in vitro. These differentiated functional cells could be used for cell therapy, disease modeling, and drug development, all valuable for research or clinical applications1,2,3. For example, a variety of methods have been developed to differentiate PSCs into cardiomyocytes (CMs)4,5,6,7. These CMs can be applied for cardiotoxicity testing of drugs, modeling of heart disease, and cell transplantation8,9,10,11.
However, the conversion from PSC to the terminal differentiated cells is a stepwise process, and multiple perturbations during the differentiation process may lead cells to divergent cell fates. Different genetic backgrounds and epigenetic marks of PSC lines influence the potential for differentiation to a specific lineage12,13,14,15; the number of PSC passages and accumulated gene mutations are also sources of PSC heterogeneity; differences in the experimental operations employed by different experimenters can also lead to completely different differentiation results16,17,18,19,20. Therefore, currently one of the main problems in PSC-derived cell production is the instability among cell lines and batches21,22,23,24,25. Instability in PSC differentiation often leads to multiple repeated experiments, consuming significant time and labor resources. To address this issue, it is crucial to develop a strategy that minimizes the variability among cell lines and batches, thus enhancing the stability and robustness of the differentiation.
Recently, advances in high-resolution microscopy and machine learning (ML) have facilitated the application of ML-based quantitative image analysis in cell biology, making it possible to utilize valuable information in cell imaging features26,27,28,29,30,31,32,33,34. In our previous work, we proposed a live-cell image-based ML strategy to monitor and intervene in the PSC differentiation status in real time to improve the stability and efficiency of the PSC differentiation (Figure 1)35. Taking PSC-to-cardiomyocyte differentiation as an example, we evaluated the initial PSC state using random forest models, predicted the optimal differentiation condition using logistic regression models, and recognized successfully differentiated cells using deep learning-based Grad-CAM36 and pix2pix37. ML models learned to identify cell lineages from a range of bright-field morphological features, including features about area, circumference, convexity, solidity, brightness, moving velocity, and other implicit features extracted by deep convolutional neural networks. Based on inference from these established ML models, we realized control of the initial PSC state, early assessment and intervention in differentiation conditions, and elimination of the misdifferentiated cell contamination, together providing a comprehensive and accurate modulation of the cardiac differentiation process. Here we provide a step-by-step protocol for developing the strategy.
Protocol
1. Cell differentiation and characterization
- Preparation of culture reagents and culture plates
- Prepare PSC culture medium by adding 2 mL of supplement and 0.2% Penicillin-Streptomycin to 48 mL of basal medium. Aliquot and store the supplement at -20 °C. Store this medium at 4 °C for up to 4 weeks.
- Prepare PSC preparation medium by adding 1 mL of supplement and 0.2% Penicillin-Streptomycin to 500 mL of basal medium. When using, preheat the medium for one-time use, and store the medium at 4 °C for up to 3 weeks.
- Prepare CM differentiation medium by adding 1x B27 supplement minus insulin and 1% Penicillin-Streptomycin into RPMI 1640. Prepare CM maintenance medium by adding 1x B27 supplement and 1% Penicillin-Streptomycin into RPMI 1640. Store these two kinds of medium at 4 °C for 2 weeks.
- Submerge the Matrigel in ice and keep it at 4 °C overnight. Ensure that Matrigel is thawed completely; then, aliquot into 1 mL aliquots stored at -80 °C for later use, and add 1 mL of Matrigel into 49 mL of DMEM/F12 at 4 °C to prepare the Matrigel working solution. Add 850 µL of this Matrigel working solution per well of a 6-well culture plate, and incubate the plate at 37 °C for at least 30 min. For the 96-well culture plate used in CM differentiation, add 35 µL of Matrigel working solution per well.
NOTE: Keep Matrigel on ice at all times; the pipette tips and tubes should be prechilled at 4 °C. Matrigel working solution could be stored at 4 °C for 2 weeks. Be careful about avoiding bubbles when pipetting the Matrigel working solution.
- PSC maintenance and passage
- Maintain PSCs in 6-well culture plates. Confirm the cell density under a microscope and prepare for passage when the confluency reaches ~70% (Figure 2A).
- Use EDTA to digest the cells for passaging. Warm PSC culture medium, PBS, and EDTA in a 37 °C water bath, and add Y27632 in the medium to a final concentration of 5 µM.
- Wash the PSCs with 1 mL of PBS, add 1 mL of EDTA per well, and incubate the plate in a 37 °C incubator for 3 min.
- Aspirate the EDTA before the cells detach from the culture plate, then use the 1 mL of PSC culture medium to gently rinse the bottom of the well 3-4x for resuspending the cells.
NOTE: This allows us to skip the step of centrifugation and reduce cell damage in passaging operations. - Aspirate the DMEM/F12 from a Matrigel-coated well of a 6-well culture plate and add 2 mL of fresh PSC culture medium per well (Y27632 added). Add the cell suspension to the wells with the split ratio of 1:6 to 1:12. Place the plate in the 37 °C incubator with 5% CO2.
- Change the PSC culture medium (without Y27632) 12 to 24 h after passaging; then, change the medium and check the cell state and confluence every day.
- Cardiac Differentiation of PSCs
- For CM differentiation, seed PSCs into a 96-well culture plate in PSC preparation medium (Y27632 added). For CM differentiation, follow the same steps above (steps 1.2.2-1.2.6) to seed PSCs into a 96-well culture plate in PSC preparation medium (Y27632 added).
NOTE: At this stage, the EDTA treatment duration could be appropriately extended (5 min or more) to sufficiently dissociate PSCs so that the clones could be smaller after passage. Our study has demonstrated that within the same area, smaller colonies are conducive to differentiation. - Stage I: When PSCs reach 80-90% confluency, switch the medium to CM differentiation medium with 2-20 µM CHIR99021 (CHIR) (Figure 2B). The time point of medium switching is referred to as day 0. After 24-48 h CHIR treatment, change the medium to fresh CM differentiation medium.
NOTE: CHIR is a GSK3beta inhibitor for mesoderm induction. The optimal CHIR dose for CM differentiation fluctuates with different cell lines and batches. For example, the optimal CHIR dose is 8-10 µM, 48 h for iPS-B1. To collect diverse bright-field images for training the ML model, here we titrated CHIR concentrations for each batch of experiments for different differentiation efficiencies. - Stage II: At 72 h (day 3), replace the medium with the CM differentiation medium supplemented with 5 µM IWR1 and culture for 48 h (day 5). Change the medium with CM differentiation medium for 1-2 days (to days 6-7). At this point, PSCs differentiate into cardiac progenitor cells (CPC).
- Stage III: Replace the medium with CM maintenance medium and replace the medium every 3 days. On day 10 or day 12, harvest the CM for further analysis.
- For CM differentiation, seed PSCs into a 96-well culture plate in PSC preparation medium (Y27632 added). For CM differentiation, follow the same steps above (steps 1.2.2-1.2.6) to seed PSCs into a 96-well culture plate in PSC preparation medium (Y27632 added).
- Immunofluorescence staining of CM
- At day 10 or day 12, wash the cells with PBS and fix them with 4% paraformaldehyde in PBS for 15 min at room temperature. Wash the CMs 2x with PBS and store the plate at 4 °C for up to ~1 week.
NOTE: Paraformaldehyde is a toxic reagent with a pungent odor. Operate in a chemical hood and protect eyes and hands. - At the time for staining, treat the cells with permeabilizing solution (0.1% Triton X-100 in PBS) for 15 min at room temperature; then, incubate cells in blocking solution (0.1% Triton X-100 containing 3% Donkey Serum in PBS) for 30 min at room temperature.
- Incubate the sample with Cardiac Troponin T (cTnT) primary antibody (1:300) diluted in blocking solution overnight at 4 °C to identify the CMs.
- Collect the primary antibody and wash the cells with PBS 3x. Incubate the sample with secondary antibodies in PBS with 1% bovine serum albumin for 1 h at 37 °C in a dark environment.
NOTE: The collected primary antibody could be reused for 2-3x in 1 week.
- Collect the primary antibody and wash the cells with PBS 3x. Incubate the sample with secondary antibodies in PBS with 1% bovine serum albumin for 1 h at 37 °C in a dark environment.
- Remove the secondary antibody from the cells and wash the cells 3x with PBS. Use Hoechst 33342 (1:1,000 in PBS) to stain nuclei for 5 min at room temperature. Rinse the cell 3x using PBS and add 100 µL of PBS per well to avoid drying. Store the plate at 4 °C in a dark place and acquire fluorescent images within 1 week.
- At day 10 or day 12, wash the cells with PBS and fix them with 4% paraformaldehyde in PBS for 15 min at room temperature. Wash the CMs 2x with PBS and store the plate at 4 °C for up to ~1 week.
2. Image stream acquisition throughout the differentiation process
- Setup of experimental design for collecting images
- Use an automated microscope supporting live cell culture and imaging to collect bright-field images of different stages in CM differentiation and cTnT immunofluorescent images of differentiation outcome (Figure 2A-E). Use the microscope-supporting software to design the experiment program and control the microscope.
- Open the software and create a new experimental design. Choose a 5x objective and a 2x tube lens for imaging. Check the Tiles and Z-stack options.
- Add TL Brightfield channel for brightfield imaging in the Channels menu. Add AF488 and H3342 channels for immunofluorescence imaging. Modify the light path in the Imaging Setup menu for imaging with the external digital CMOS camera.
- Open the Acquisition Mode menu, and set 2 x 2 binning to increase the signal-to-noise ratio.
- Open the Z-stack menu and choose the Center model. Set the number of slices and intervals when scanning. Choose 3-5 slices with 3-6 µm intervals for in-focus images in analysis.
- In the Navigation & Tiles window, set up the tile regions by Carrier and set 25 tiles (5 columns x 5 rows) for one well. In the Tiles menu, select the Multiwell 96 template in Sample Carrier, and set an overlap of 5%-15% on adjacent regions.
- If continuous imaging is needed, check the Time series option and set the duration and interval in the Time series menu. For collecting the image stream at stage I, set a 12 h duration without intervals.
- Acquisition of bright-field and immunofluorescent images
- For acquiring images, first put the cell culture plate into the sample tray, and load the sample inside the microscope. Select the Multiwell 96 template to match the experimental design. If the sample consists of live cells, open the heating system and CO2 pump to keep the appropriate condition for culture (37 °C, 5% CO2).
- Open the preset experimental project and save it as a new one. Open the Tiles menu and calibrate the position of the plate manually. In the Navigation & Tiles window, select the wells needed and click Create to construct tile regions for these wells.
- Click the Verify Tile Regions in the Tiles menu and run Autofocus to verify all of the wells. Then, manually correct the focusing of each well under the brightfield by scrolling the mouse wheel.
- Tick the channels needed and set the times of exposure in the Channels menu. The recommended exposure times are 1.625 ms for Bright, 100 ms for H3342, and 550 ms for AF488. Execute a single snap to check the images of all channels.
- Finally, click the Start experiment button, and wait for automatic imaging. Typically, it takes ~1.2 h to finish the scanning of a whole 96-well culture plate (96 wells, three layers, 25 tiles for each well under the brightfield). In the Processing framework, choose Image Export, select the file type of the uncompressed TIFF format or PNG format, and apply. Use images of 9 tiles (3 x 3) in the center of well for the further analysis (Figure 2A-E).
3. Establishment of the image-based ML strategy at each stage of the differentiation process
- Software preparation
- Install the Python environment, with packages pytorch (1.9.0), numpy, scipy, scikit-learn, pandas, visdom, scikit-image, opencv-python, and matplotlib.
- Install Jupyter Notebook.
- Install MATLAB.
- Download the code and the example datasets from https://github.com/zhaoyanglab/ML-for-PSC-differentiation.
NOTE: The performance of ML is dependent on the quality of datasets and the choice of models and hyper-parameters. The example implementation is not guaranteed to be optimal for all circumstances. ML engineers should customize the code and tailor the model and hyper-parameters for their specific needs.
- ML strategy at the PSC stage: Control of initial PSC colony states
Dataset: pairs of brightfield images at 0 h (before CHIR treatment) and final differentiation efficiency
Model: random forest- Prepare a dataset consisting of brightfield images at 0 h (before CHIR treatment) and the final cTnT fluorescence images.To increase the diversity of morphological profiles of 0 h brightfield images, ensure that the dataset includes PSC colonies from different cell lines, with differentiation initiated over a time interval after passage. Treat wells in the dataset with the optimal CHIR conditions at stage I.
NOTE: To ensure that colonies are treated with the optimal CHIR conditions, wells within the same batch can be treated with titrated CHIR doses. Only wells with high-efficiency CHIR conditions are included in the dataset. - Quantify the differentiation efficiency of each well by the Differentiation Efficiency Index computed from its cTnT fluorescence images, which is defined by
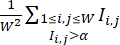 (where
(where 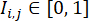 represents the fluorescence intensity at pixel location (i, j) in the W × W fluorescence image, and the sum is taken over all pixel locations with intensity above a threshold α, typically set at 0.5). Within each cell line, compute the maximum Differentiation Efficiency Index over all the wells in this cell line, and normalize their Differentiation Efficiency Indexes by the maximum value.
represents the fluorescence intensity at pixel location (i, j) in the W × W fluorescence image, and the sum is taken over all pixel locations with intensity above a threshold α, typically set at 0.5). Within each cell line, compute the maximum Differentiation Efficiency Index over all the wells in this cell line, and normalize their Differentiation Efficiency Indexes by the maximum value.
NOTE: Since the differentiation potential of each cell line is inherently different, such a normalization will emphasize more on the colony morphological features shared among various cell lines that contribute to the differentiation outcome. Normalization indeed makes PSC colonies from different cell lines comparable and mitigates the bias induced by cell lines. - Quantify the morphological profiles of 0 h brightfield images by high-dimensional features that reveal the properties of the colony shape. To outline colony regions in brightfield images, compute the local entropy (with radius r); pixels with local entropy greater than a threshold σ are identified by covered by cells. Use a parameter setting of r = 10 pixels and σ = 3. Switch the working directory to "./colony_control"; save the brightfield images, binary images of cell regions, and cTnT fluorescence images at "colony_control/ image_data/CD00-*/[brightfield|cell_region|ctnt]/S*.png"; and run the Jupyter Notebook "Compute_features.ipynb" to compute features related to colony area, circumference, solidity, convexity, circularity, and centroid-contour distances, among others.
- Randomly divide the dataset into a training set and a test set, with the training set comprising between 50% and 80% of the overall data. Run the final block Split the dataset in the Jupyter Notebook to split the dataset (Figure 3A).
- Train a random forest regression model on the training set to predict differentiation efficiency from 0 h brightfield image features. Run the Jupyter Notebook "Machine_learning.ipynb" to train and test the model.
- Evaluate the trained random forest model on the test set. Confirm that the predicted differentiation efficiency highly correlates with the true one using the Pearson correlation coefficient (Figure 3B).
NOTE: Since the test set is not involved in the training phase, a high consistency between the predicted and the true differentiation efficiency on the test set indicates that the model can generalize well on unseen samples. - Obtain feature importance from the trained random forest model and select 8 features with the most importance. Visualize their relationship with the final differentiation efficiency by dividing the range of features into bins and computing the mean differentiation efficiency in each bin. Run the Jupyter Notebook "Feature_importance.ipynb" to perform the visualization (Figure 3C).
NOTE: The visualization may help understand how changes in PSC colony shape contribute to the differentiation. - Apply the trained model in new batches to monitor PSC colony states in real time. Capture the bright-field image at every hour, pass the image to the trained model, and obtain the predicted differentiation efficiency. Once the predicted efficiency exceeds a threshold (e.g., 50%), start the differentiation process. If the predicted efficiency could not reach the threshold, discard these samples.
NOTE: During the PSC stage, the differentiation potential of PSC colonies will first rise and later decline. Real-time monitoring enables experimenters to identify the most conducive time point to initiate differentiation.
- Prepare a dataset consisting of brightfield images at 0 h (before CHIR treatment) and the final cTnT fluorescence images.To increase the diversity of morphological profiles of 0 h brightfield images, ensure that the dataset includes PSC colonies from different cell lines, with differentiation initiated over a time interval after passage. Treat wells in the dataset with the optimal CHIR conditions at stage I.
- ML strategy at stage I: early assessment and adjustment of CHIR doses
Dataset: pairs of brightfield image streams at 0-12 h and CHIR concentration labels (low/optimal/high)
Model: logistic regression- Prepare a dataset consisting of whole-well brightfield image streams, where images are acquired every hour between 0 h and 12 h. Record the CHIR doses (i.e., combinations of CHIR concentrations and duration) of each well. Ensure that the dataset includes different batches and wells with titrated CHIR doses (e.g., concentration = 2, 4, 6, 8, 10, and 12 µM; duration = 24, 36, and 48 h) to increase the diversity. Switch the working directory to "./stage_I"; save the images as "./data/image/CD01-*/S*/T*.png", where CD01-* (CD01-1, CD01-2, CD01-3, CD01-4) is the batch name, S* (S1, S2, ..., S96) is the index of well, and T*.png (T1.png, T2.png, ..., T10.png) is the preprocessed bright-field image of the well at different time step of the image stream.
- In each batch, compute the percentage of cTnT+ cells for each well, which is calculated by dividing the number of cTnT+ cells by the total number of Hoechst+ cells in the fluorescence image. Average the percentage of cTnT+ cells over parallel wells with the same CHIR dose condition. Save the CHIR dose condition and percentage of cTnT+ cells for each well in a csv file "./data/image/CD01-*/CD01-?_exp_config.csv".
- In each batch, determine the "low", "optimal", and "high" CHIR concentration range under each CHIR duration according to the following criteria. CHIR concentrations with an average percentage of cTnT+ cells above a threshold (e.g., 20%) are identified as "optimal". CHIR concentrations beyond the optimal range are identified as "low" or "high". Under each CHIR duration, label the wells as "low", "optimal", and "high" according to their corresponding CHIR concentrations.
NOTE: Empirically, there is a negative correlation between CHIR concentrations and the duration of high-efficiency wells within a batch: for higher CHIR duration, the optimal concentrations decrease. In other words, the optimality of CHIR concentrations depends on CHIR duration, and thus, the labels of wells differ among different CHIR durations. - In each batch, under each CHIR duration, compute ΔCHIR concentration for each concentration c to quantify its deviation from optimum. Denote the optimal CHIR concentration range by [c1, c2], and then ΔCHIR concentration(c) = c - c1 (if c < c1), 0 (if c1 ≤ c ≤ c2), or c - c2 (if c > c2).
- Extract features of the image streams in the dataset, ensuring that they reflect the morphological changes of cells during stage I. To compute image features about local entropy, area, circumferences, brightness, optical flow, and fractal dimension, run
cd ./data
python compute_features.py
cd ..
NOTE: To highlight the morphological changes, the values of some features (e.g., area, circumferences, and optical flow) can be normalized by their initial baseline values in the image streams. - Randomly divide the dataset into a training set and a test set. Run the Jupyter Notebook "./data/prepare_dataset.ipynb" to split the dataset and organize the datasets as Dataframes (Figure 3D).
- For each CHIR duration, train a logistic regression model to predict the CHIR concentration label (low/optimal/high) from the image stream features on the training set. Evaluate the classification performance of the trained logistic regression model on the test set using accuracy, precision, recall, F1 score, and area under the curve (AUC).Run the Jupyter Notebook "machine_learning.ipynb" for training and testing (Figure 3E).
- Evaluate the model performance in CHIR dose assessment. In the test set, merge predicted labels of parallel wells with the same CHIR concentration using Deviation Scores (ranging from -1 to 1), defined by
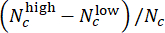 , where Nc denotes the number of wells with concentration c and
, where Nc denotes the number of wells with concentration c and 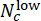 and
and 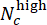 wells among them are predicted as "low" and "high", respectively. Confirm that the predicted Deviation Scores highly correlate with the true ΔCHIR concentration for each CHIR dose using the Pearson correlation coefficient.
wells among them are predicted as "low" and "high", respectively. Confirm that the predicted Deviation Scores highly correlate with the true ΔCHIR concentration for each CHIR dose using the Pearson correlation coefficient. - Perform a cross-batch validation to test the model's generalization performances. On each round, leave one batch for testing while using others for training. Compare the predicted Deviation Scores and true ΔCHIR concentration for each CHIR dose in every test batch using the Pearson correlation coefficient. Run the Jupyter Notebook "cross_batch_validation.ipynb" to implement the cross-batch validation (Figure 3F).
NOTE: For a selected CHIR duration h, prediction for all the wells with concentration from the logistic regression model trained under duration h will be converted to a Deviation Score, reflecting the optimality of CHIR concentration c. A Deviation Score close to 0 suggests that concentration c is predicted to be optimal under duration h, while a negative or positive Deviation Score suggests an underdose or overdose condition, which can serve as a signal for later adjustment of CHIR doses. Note that since the prediction is made early (12 h), the final CHIR duration and CHIR concentration during the rest of stage I can still be adjusted. - Apply the trained logistic regression models to assess CHIR doses in new batches. For a given CHIR concentration , compute 0-12 h image stream features for parallel wells with concentration , pass the features to the logistic regression models trained under different CHIR durations, and accordingly, obtain the predicted labels for each well. Based on the models' prediction, compute Deviation Scores to assess each CHIR concentration under different CHIR durations.
- With the model-based CHIR dose assessment, rescue wells under each suboptimal CHIR concentration accordingly, by adjusting their CHIR duration or concentration towards optimum before 48 h.
- Adjusting duration: according to the model prediction about the optimal CHIR duration, adjust the time point when the CHIR-containing medium in these wells is switched to the CM differentiation medium. If the optimal time point is missed, discard these samples.
- Adjusting concentration: under the preset CHIR duration, if the current concentration is predicted as "low", directly add a high concentration of CHIR into the medium to increase CHIR concentration. Dilute CHIR in DMSO (50 mM) with PBS when needed, add an appropriate volume of liquid, and gently shake the plate. If the current concentration is predicted as "high", aspirate a certain volume of CHIR-containing medium and supplement with the same volume of CM differentiation medium.
NOTE: In the first 48 h of differentiation, cells will secrete factors to improve cell growth and differentiation, so do not completely change the medium. In addition, it is necessary to keep the medium volume constant in differentiation. Therefore, make sure the volume of liquid added is as small as possible when adjusting CHIR concentration.
- ML strategy at stage II: recognition of CM-committed CPCs
Dataset: pairs of bright-field images on day 6 and manually annotated masks of CM-committed CPCs
Model: ResNeSt38 with Grad-CAM36 for weakly supervised learning- Prepare a dataset consisting of brightfield images on day 6. Manually annotate CM-committed CPCs in the brightfield images by tracking the cTnT+ cells in the image streams from day 12 back to day 6.
- Crop the brightfield images and the corresponding manual annotation (termed masks) of CM-committed CPCs into patches, with each patch sized at 1/5.5 of the whole-well image size. Extract patches in an overlapping manner where adjacent patches share a 50% overlap in the training set and a 75% overlap in the test set. Label patches with ≥30% CM-committed CPCs as "positive". Label patches without CM-committed CPCs as "negative". Switch the working directory to "stage_II". Run "./Crop_and_Reconstruct/mask_crop.m" to crop the images. Run "./Crop_and_Reconstruct/classification.m" to assign labels (0: negative; 1: positive) to brightfield patches (Figure 3G).
- Randomly divide the labeled image patches into a training set and a test set.
- Train a deep convolutional neural network, ResNeSt38 to learn to classify these patches. Evaluate the trained ResNeSt on the test set to confirm the model demonstrates a high classification accuracy. Run "train.py" and "test.py" for training and testing, respectively (Figure 3H).
- Use Grad-CAM36 to highlight the regions that contribute most to the inference of ResNeSt, represented by heatmaps. Binarize the heatmaps (using a threshold, e.g., 10% × maximum) to get the predicted CM-committed CPC regions, which are named imaged-recognized CPC (IR-CPC) regions.
- Compare the IR-CPC regions with manually annotated masks on the test set using accuracy, F1 score, precision, recall, specificity, and intersection-over-union (IoU). Compare the proportion of IR-CPC regions with Differentiation Efficiency Indexes (computed from the final cTnT fluorescence images) using the Pearson correlation coefficient. Run "./Evaluation/index_calculate.m" and "./Evaluation/Pearson_correlation.m" for performance evaluation (Figure 3I).
- Apply the trained model to recognize CPCs for new batches at stage II. Crop whole-well brightfield images into patches, pass the patches to the trained ResNeSt, and obtain the IR-CPC regions at the patch level, which are then merged to yield the whole-well IR-CPC regions. Compute the fraction of IR-CPC regions as an indicator of the final differentiation efficiency. Purify the IR-CPCs to reduce cell contamination (described below).
NOTE: Since the CPC recognition model can make an early assessment of the final differentiation efficiency, it can also serve as a high-throughput readout for screening small molecules that are conducive to the differentiation process, which may further optimize the differentiation protocol35.
- Region-selective purification of imaged-recognized CPCs
NOTE: Photoactivation experiments are based on an inverted fluorescence microscope facilitated with a motorized stage.- According to the IR-CPC regions predicted by the trained model, discard the wells that could not generate any IR-CPCs. To purify these CM-committed CPCs, use a non-cytotoxic photoactivatable probe, Dual-Activatable Cell Tracker 1 (DACT-1) to region-selectively label non-CPC39.
- Dissolve the DACT-1 (10 mM) in DMSO and store it at -20 °C. Incubate day 6 cells with 1 µM DACT-1 in CM differentiation medium for 30 min at 37 °C.
- Inspect the live cells under a microscope and select the region of interest (ROI) as polygons in MATLAB. Draw parallel lines with 20 µm spacing inside the polygon and calculate the coordinates of the intersection points.
- Set the moving track of the motorized stage according to trace lines, then the stage would move at 0.12 mm/s so that a 405nm laser line (20 µm in diameter) assembled on the stage could globally scan the ROI to photoactivate non-CPCs restrictedly. After irradiation, detect the DACT-1-labeled cells using a 561 nm laser line.
- Dissociate cells by using 0.05% Trypsin-EDTA (diluted by PBS) at 37 °C for 5-7 min, and gently shake the plate in an incubator for 2 min. Filter the cells through a 40 µm cell strainer and centrifuge at 500 × g for 3 min. Resuspend the cells in 0.5% BSA and keep the tube on ice until sorting. Also prepare an unirradiated sample as the negative control for sorting.
- Start up the flow cytometry sorting system and set up the sorter software. Choose the 561 nm wavelength lasers and use a bandpass filter of 610/20 for DACT1+ sorting. For each sample, use a polygonal gating tool to circle the cell population in a bivariate plot of forward scatter (FSC) and side scatter (SSC) for selecting the single cell. Then, run the negative control to confirm the background fluorescence, run the region-selectively irradiated sample, and draw the gate for the DACT1+ population.
- Collection and resuspend the sorted cells in CM maintenance medium with 10% FBS and 5 µM Y27632, and seed the sorted cells in a Matrigel-coated 96-well plate.
- The next day, change the medium to CM maintenance medium and wait to harvest high-purity CMs.
- ML strategy at stage III: recognition of CMs
Dataset: pairs of brightfield images on day 12 and the final cTnT fluorescence images
Model: pix2pix37- Prepare a dataset consisting of brightfield images of CMs and the corresponding cTnT fluorescence images. To increase the diversity, ensure that the dataset includes wells with various differentiation efficiencies and cell lines.
- Randomly divide the dataset into a training set and a test set. Switch the working directory to "stage_III". Save the brightfield images and the corresponding cTnT fluorescence images at "./pix2pix/datasets/(CM|CM_new_cell_lines)/A/(train|test)/*.png" and ".../B/(train|test)/*.png", respectively. Run the following command:
cd pix2pix/datasets
python combine_A_and_B.py --fold_A ./CM/A --fold_B ./CM/B --fold_AB ./CM/ --no_multiprocessing
cd .. - Train the pix2pix model37 on the training set. Run the following command (Figure 3J):
python train.py --dataroot ./datasets/CM --name brightfield2fluorescence --model pix2pix --input_nc 1 --output_nc 1 --load_size 1536 --crop_size 256 --lr 2e-4 --n_epochs 1000 --n_epochs_decay 0 --norm instance --netD n_layers --n_layers_D 1 --batch_size 16 --direction AtoB --save_epoch_freq 100 --dataset_mode aligned --use_resize_conv --seed 1234 - Apply the trained pix2pix model on the test set.Run the following command:
python test.py --dataroot ./datasets/CM --name brightfield2fluorescence --model pix2pix --direction AtoB --input_nc 1 --output_nc 1 --load_size 1536 --crop_size 1536 --use_resize_conv --eval --num_test 1000 - Compare the fluorescence images predicted by the model with the true (i.e., experimentally obtained) ones. At the pixel level, compare the predicted intensities and true intensities using the Pearson correlation coefficient. In addition, at the image level, compare predicted Differentiation Efficiency Indexes with the true ones using the Pearson correlation coefficient. Run "stage_III/evaluation/pixel_correlation.m" and "image_correlation.ipynb" for these comparisons (Figure 3K,L).
- Apply the trained model to recognize CMs in new batches at stage III. Pass new brightfield images of CMs to the trained pix2pix model and obtain an in-silico prediction of the fluorescence images and differentiation efficiency.
NOTE: To reduce cell contamination, DACT-1-based photoactivated cell sorting is also applicable for purifying CMs recognized by the model.
Results
Based on brightfield imaging and ML, the overall differentiation process can be intelligently monitored and optimized. At the PSC stage, we developed an ML model that could predict the final differentiation efficiency according to the morphological features of initial PSC colonies, to determine the most suitable or appropriate time point to initiate differentiation (Figure 4A,B). The differentiation efficiency predicted by the random forest model is highly correlated w...
Discussion
Here, we described a detailed protocol to overcome one of the major problems in current PSC application and translation—the variability in cell differentiation. By harnessing live-cell brightfield imaging and ML, we iteratively optimized PSC differentiation to achieve consistently high efficiency across cell lines and batches. However, in the above differentiation process, several critical steps in the protocol have a decisive influence on whether the differentiation would succeed or not. Since the cell state in th...
Disclosures
Yang Zhao, Jue Zhang, Xiaochun Yang, Yao Wang, and Daichao Chen are filing a patent for the PSC differentiation strategy reported in this paper (202210525166.X).
Acknowledgements
We thank Qiushi Sun, Yao Wang, Yu Xia, Jinyu Yang, Chang Lin, Zimu Cen, Dongdong Liang, Rong Wei, Ze Xu, Guangyin Xi, Gang Xue, Can Ye, Li-Peng Wang, Peng Zou, Shi-Qiang Wang, Pablo Rivera-Fuentes, Salome Püntener, Zhixing Chen, Yi Liu, and Jue Zhang, for laying the groundwork of this strategy. This work was supported by the National Key R&D Program of China (2018YFA0800504, 2019YFA0110000) and the Space Medical Experiment Project of China Manned Space Program (HYZHXM01020) to Yang Zhao. Figure 1 was created with BioRender.com.
Materials
| Name | Company | Catalog Number | Comments |
| 0.25% Trypsin-EDTA | Gibco | 25200056 | Diluted digests were used for CPC and CM digestion |
| 4% Paraformaldehyde in PBS | KeyGEN BioTECH | KGIHC016 | |
| 6-well Cell Culture Plate | NEST | 703001 | |
| 96-well Cell Culture Plate | NEST | 701001 | |
| B27 Supplement | Gibco | 17504044 | |
| B27 Supplement Minus Insulin | Gibco | A1895601 | |
| Bovine serum albumin (BSA) | GPC BIOTECH | AA904-100G | |
| Celldiscoverer 7 | Zeiss | Instruments used to take bright-field images throughout differentiation and final cTnT images | |
| CHIR99021 | Selleck | S1263 | |
| DMEM/F12 | Gibco | 12634010 | |
| Donkey anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor 488 | Thermo | A-21202 | Secondary Antibody |
| FACSAria III | BD Biosciences | Flow cytometry sorter | |
| Fetal Bovine Serum (FBS) | VISTECH | SE100-B | |
| Hoechst 33342 | YEASEN | 40732ES03 | |
| Human Pluripotent Stem Cell Chemical-defined Medium | Cauliscell Inc | 400105 | Basal medium of PSC preparation medium |
| iPS-18 | TaKaRa | Y00300 | |
| iPS-B1 | Cellapy | CA4025106 | |
| iPS-F | Nuwacell | RC01001-B | |
| iPS-M | Nuwacell | RC01001-A | |
| IWR1-1-endo | Selleck | S7086 | IWR1 |
| Jupyter Notebook | N/A | Version 6.4.0 | https://jupyter.org/ |
| MATLAB | MathWorks | Version R2020a | Software for scientific computation and image annotation |
| Matrigel Matrix | Corning | 354230 | Matrigel |
| Mouse monoclonal IgG1 anti-cTnT | Thermo | MA5-12960 | cTnT primary antibody |
| Normal Donkey Serum | Jackson | 017-000-121 | |
| ORCA-Flash 4.0 V3 digital CMOS camera | Hamamatsu | C13440-20CU | The digital camera assembled on Celldiscoverer7 |
| PBS | NEB | 21-040-CVR | |
| Penicillin-Streptomycin | Gibco | 15140-122 | |
| Pluripotency Growth Mater 1 basal medium | Cellapy | CA1007500-1 | Basal medium of PSC culture medium |
| Pluripotency Growth Mater 1 supplement | Cellapy | CA1007500-2 | Supplement of PSC culture medium |
| Prism | Graphpad | Version 8/9 | Statistical software for statistical analysis and plotting |
| Python | N/A | version 3.6 | Python 3 environment for scientific computation, with packages pytorch (1.9.0), numpy, scipy, pandas, visdom, scikit-learn, scikit-image, opencv-python, and matplotlib software for scientific computation and image annotation. |
| RPMI 1640 | Gibco | 11875176 | |
| Supplement hPSC-CDM (500x) | Cauliscell Inc | 00015 | Supplement of PSC preparation medium |
| TiE | Nikon | An inverted fluorescence microscope (with modification) for region-selevtive purification | |
| Triton X-100 | Amresco | 9002-93-1 | |
| Versene Solution | Thermo | 15040066 | EDTA solution for PSC digestion |
| Y27632 | Selleck | S6390 | |
| Zen | Zeiss | Version 3.1 | A supporting software of Celldiscoverer7 for image acquisition, processing and analysis |
References
- Yoshida, Y., Yamanaka, S. Induced pluripotent stem cells 10 years later: for cardiac applications. Circ Res. 120 (12), 1958-1968 (2017).
- Shi, Y., Inoue, H., Wu, J. C., Yamanaka, S. Induced pluripotent stem cell technology: a decade of progress. Nat Rev Drug Discov. 16 (2), 115-130 (2017).
- Takahashi, K., et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 131 (5), 861-872 (2007).
- Burridge, P. W., et al. Chemically defined generation of human cardiomyocytes. Nat Methods. 11 (8), 855-860 (2014).
- Mordwinkin, N. M., Burridge, P. W., Wu, J. C. A review of human pluripotent stem cell-derived cardiomyocytes for high-throughput drug discovery, cardiotoxicity screening, and publication standards. J Cardiovasc Transl Res. 6 (1), 22-30 (2013).
- Vanden Berg, C. W., Elliott, D. A., Braam, S. R., Mummery, C. L., Davis, R. P. Differentiation of human pluripotent stem cells to cardiomyocytes under defined conditions. Methods Mol Biol. 1353, 163-180 (2016).
- Burridge, P. W., Holmström, A., Wu, J. C. Chemically defined culture and cardiomyocyte differentiation of human pluripotent stem cells. Curr Protoc Hum Genet. 87, 1-15 (2015).
- Li, J., et al. Human pluripotent stem cell-derived cardiac tissue-like constructs for repairing the infarcted myocardium. Stem Cell Rep. 9 (5), 1546-1559 (2017).
- Smith, A. S. T., Macadangdang, J., Leung, W., Laflamme, M. A., Kim, D. -. H. Human iPSC-derived cardiomyocytes and tissue engineering strategies for disease modeling and drug screening. Biotechnol Adv. 35 (1), 77-94 (2017).
- Sharma, A., et al. Use of human induced pluripotent stem cell-derived cardiomyocytes to assess drug cardiotoxicity. Nat Protoc. 13 (12), 3018-3041 (2018).
- Mehta, A., et al. Identification of a targeted and testable antiarrhythmic therapy for long-QT syndrome type 2 using a patient-specific cellular model. Eur Heart J. 39 (16), 1446-1455 (2018).
- Cahan, P., Daley, G. Q. Origins and implications of pluripotent stem cell variability and heterogeneity. Nat Rev Mol Cell Biol. 14 (6), 357-368 (2013).
- Kim, K., et al. Epigenetic memory in induced pluripotent stem cells. Nature. 467 (7313), 285-290 (2010).
- Rouhani, F., et al. Genetic background drives transcriptional variation in human induced pluripotent stem cells. PLoS Genet. 10 (6), e1004432 (2014).
- Kim, K., et al. Donor cell type can influence the epigenome and differentiation potential of human induced pluripotent stem cells. Nat Biotechnol. 29 (12), 1117-1119 (2011).
- Garitaonandia, I., et al. Increased risk of genetic and epigenetic instability in human embryonic stem cells associated with specific culture conditions. PLoS One. 10 (2), e0118307 (2015).
- Engler, A. J., Sen, S., Sweeney, H. L., Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell. 126 (4), 677-689 (2006).
- Matoba, N., Yamashita, T., Takayama, K., Sakurai, F., Mizuguchi, H. Optimal human iPS cell culture method for efficient hepatic differentiation. Differentiation. 104, 13-21 (2018).
- Volpato, V., et al. Reproducibility of molecular phenotypes after long-term differentiation to human iPSC-derived neurons: A multi-site omics study. Stem Cell Rep. 11 (4), 897-911 (2018).
- Anderson, N. C., et al. Balancing serendipity and reproducibility: Pluripotent stem cells as experimental systems for intellectual and developmental disorders. Stem Cell Rep. 16 (6), 1446-1457 (2021).
- Sepac, A., et al. Comparison of cardiomyogenic potential among human ESC and iPSC lines. Cell Transplant. 21 (11), 2523-2530 (2012).
- Laco, F., et al. Unraveling the inconsistencies of cardiac differentiation efficiency induced by the GSK3β inhibitor CHIR99021 in human pluripotent stem cells. Stem Cell Rep. 10 (6), 1851-1866 (2018).
- Volpato, V., Webber, C. Addressing variability in iPSC-derived models of human disease: guidelines to promote reproducibility. Dis Model Mech. 13 (1), 042317 (2020).
- Hu, B. -. Y., et al. Neural differentiation of human induced pluripotent stem cells follows developmental principles but with variable potency. Proc Natl Acad Sci USA. 107 (9), 4335-4340 (2010).
- Huo, J., et al. Evaluation of batch variations in induced pluripotent stem cell-derived human cardiomyocytes from 2 major suppliers. Toxicol Sci. 156 (1), 25-38 (2017).
- Rostam, H. M., Reynolds, P. M., Alexander, M. R., Gadegaard, N., Ghaemmaghami, A. M. Image based machine learning for identification of macrophage subsets. Sci Rep. 7 (1), 3521 (2017).
- Christiansen, E. M., et al. In silico labeling: Predicting fluorescent labels in unlabeled images. Cell. 173 (3), 792-803 (2018).
- Buggenthin, F., et al. Prospective identification of hematopoietic lineage choice by deep learning. Nat Methods. 14 (4), 403-406 (2017).
- Smith, Z. D., Nachman, I., Regev, A., Meissner, A. Dynamic single-cell imaging of direct reprogramming reveals an early specifying event. Nat Biotechnol. 28 (5), 521-526 (2010).
- Fan, K., Zhang, S., Zhang, Y., Lu, J., Holcombe, M., Zhang, X. A machine learning assisted, label-free, non-invasive approach for somatic reprogramming in induced pluripotent stem cell colony formation detection and prediction. Sci Rep. 7 (1), 13496 (2017).
- Kusumoto, D., et al. Automated deep learning-based system to identify endothelial cells derived from induced pluripotent stem cells. Stem Cell Rep. 10 (6), 1687-1695 (2018).
- Zhu, Y., et al. Deep learning-based predictive identification of neural stem cell differentiation. Nat Commun. 12 (1), 2614 (2021).
- Qian, T., et al. Label-free imaging for quality control of cardiomyocyte differentiation. Nat Commun. 12 (1), 4580 (2021).
- Ounkomol, C., Seshamani, S., Maleckar, M. M., Collman, F., Johnson, G. R. Label-free prediction of three-dimensional fluorescence images from transmitted-light microscopy. Nat Methods. 15 (11), 917-920 (2018).
- Yang, X., et al. A live-cell image-based machine learning strategy for reducing variability in PSC differentiation systems. Cell Discovery. 9 (1), 53 (2023).
- Selvaraju, R. R., et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. Proc IEEE Int Conf Computer Vision (ICCV). , 618-626 (2017).
- Isola, P., Zhu, J. -. Y., Zhou, T., Efros, A. A. Image-to-image translation with conditional adversarial networks. Proc IEEE Conf Computer Vision Pattern Recognition (CVPR). , 5967-5976 (2017).
- Zhang, H., et al. ResNeSt: Split-attention networks. Proc IEEE/CVF Conf Computer Vision Pattern Recognition Workshops (CVPRW). , 2735-2745 (2022).
- Halabi, E. A., et al. Dual-activatable cell tracker for controlled and prolonged single-cell labeling). ACS Chem. Biol. 15 (6), 1613-1620 (2020).
- Hannan, N. R. F., Segeritz, C. -. P., Touboul, T., Vallier, L. Production of hepatocyte-like cells from human pluripotent stem cells. Nat Protoc. 8 (2), 430-437 (2013).
- Zhang, H., Shen, M., Wu, J. C. Generation of quiescent cardiac fibroblasts derived from human induced pluripotent stem cells. Methods Mol Biol. 2454, 109-115 (2022).
- Harding, M. J., McGraw, H. F., Nechiporuk, A. The roles and regulation of multicellular rosette structures during morphogenesis. Development. 141 (13), 2549-2558 (2014).
- Najm, F. J., et al. Rapid and robust generation of functional oligodendrocyte progenitor cells from epiblast stem cells. Nat Methods. 8 (11), 957-962 (2011).
- Hogrebe, N. J., Maxwell, K. G., Augsornworawat, P., Millman, J. R. Generation of insulin-producing pancreatic β cells from multiple human stem cell lines. Nat Protoc. 16 (9), 4109-4143 (2021).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved