A subscription to JoVE is required to view this content. Sign in or start your free trial.
Development and Application of Rapamycin-regulated Tyrosine Phosphatases
In This Article
Summary
This protocol describes the design, creation, and application of rapamycin-regulated phosphatases. This method provides high specificity and tight temporal control of phosphatase activation in living cells.
Abstract
Tyrosine phosphatases are an important family of enzymes that regulate critical physiological functions. They are often dysregulated in human diseases, making them key targets of biological studies. Tools that enable the regulation of phosphatase activity are instrumental in the dissection of their function. Traditional approaches, such as overexpression of constitutively active or dominant negative mutants, or downregulation using siRNA, lack temporal control. Phosphatase inhibitors often have poor specificity, and they only allow researchers to determine what processes are affected by the inhibition of the phosphatase.
We developed a chemogenetic approach, the Rapamycin-regulated (RapR) system, which allows for allosteric regulation of a phosphatase catalytic domain that enables tight temporal control of phosphatase activation. The RapR system consists of an iFKBP domain inserted into an allosteric site in the phosphatase. The intrinsic structural dynamics of the RapR domain disrupt the catalytic domain, leading to the inactivation of the enzyme. The addition of rapamycin mediates the formation of a complex between iFKBP and a co-expressed FRB protein, which stabilizes iFKBP and restores activity to the phosphatase's catalytic domain.
This system provides high specificity and tight temporal control of phosphatase activation in living cells. The unique capabilities of this system enable the identification of transient events and interrogation of individual signaling pathways downstream of a phosphatase. This protocol describes guidelines for the development of a RapR-phosphatase, its biochemical characterization, and the analysis of its effects on downstream signaling and regulation of cell morphodynamics. It also provides a detailed description of a protein engineering strategy, in vitro assays analyzing phosphatase activity, and live cell imaging experiments identifying changes in cell morphology.
Introduction
Protein tyrosine phosphatases are a critical family of proteins involved in a plethora of cell signaling events. They have been shown to play a key role in the regulation of cell proliferation, migration, and apoptosis1,2,3. Consequently, the dysregulation of protein tyrosine phosphatases leads to a variety of debilitating diseases and disorders4,5,6,7. Studying the physiological function of tyrosine phosphatases and their role in the development of these pathologies has been historically hindered by a lack of tools needed for probing the intricacies of phosphatase signaling8.
Traditionally, phosphatases are studied using methods that do not have the desired specificity and/or do not provide temporal control of their activity. These critical limitations of available tools make it challenging to dissect specific roles of phosphatases in signaling pathways. Overexpression of constitutively active and dominant negative mutants or downregulation of expression of the phosphatase provide specificity but lack temporal control and often can trigger compensatory mechanisms that will mask the true function of the enzyme.
Pharmacological inhibitors allow for the temporal regulation of phosphatases. However, many phosphatase inhibitors are notoriously nonspecific due to the well-conserved composition of the active site in tyrosine phosphatases9. Additionally, due to design constraints, inhibitors targeting the catalytic site exhibit poor cell membrane permeability10. Another limitation of inhibitors is that they only allow for the examination of the effects of phosphatase inactivation11. Thus, there is a need for tools that enable specific, temporally regulatable activation of phosphatases. These tools will allow researchers to identify the direct effects of phosphatase activation, separating them from multiple parallel signaling cascades often activated by biological stimuli. Importantly, tight temporal control of activity enables the identification of transient events induced by a phosphatase and separates the effects of acute and prolonged phosphatase activity. Combining temporal regulation with mutational analysis will allow for a detailed dissection of specific roles of individual domains of the phosphatase and interrogation of its downstream signaling12.
To address the lack of desired capabilities in the existing tools, the Karginov group has developed the Rapamycin Regulated (RapR) system13,14,15. The RapR system utilizes an engineered switch domain, iFKBP, that allows for allosteric regulation of the protein of interest (POI). The insertion of the iFKBP domain at a position allosterically coupled to the catalytic site of the POI renders it susceptible to regulation by rapamycin. In the absence of rapamycin, iFKBP disrupts the catalytic site due to the intrinsically high structural dynamics of iFKBP and thus inactivates the POI. The addition of rapamycin induces the interaction of iFKBP with the co-expressed protein FRB (Figure 1). This causes stabilization of the switch domain, which consequently restores the structure and function of the POI's catalytic domain. As such, the tool allows for specific and temporally regulatable activation of the POI.
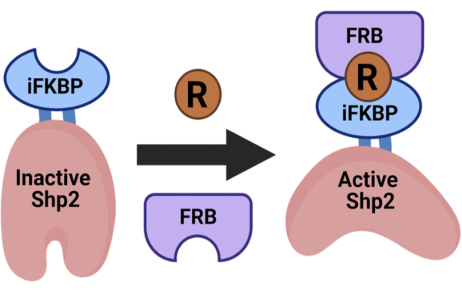
Figure 1: Schematic of the RapR-Shp2 rapamycin-regulated system. RapR allows for allosteric activation of the protein of interest with the addition of rapamycin. This figure was modified from Fauser et al.12. Abbreviations: iFKBP = insertable FKBP12; FRB = FKBP-rapamycin-binding domain; R = rapamycin; Shp2 = Src homology-2 domain-containing protein tyrosine phosphatase. Please click here to view a larger version of this figure.
The RapR tool can be applied to different protein families. It can be used to regulate protein kinases as well as phosphatases12,14. This protocol will focus on the application of the RapR tool to control Shp2 phosphatase. Shp2 is a ubiquitously expressed protein tyrosine phosphatase that is involved in signaling processes such as proliferation, migration, immunomodulation, and differentiation1,16,17,18. Dysregulation of Shp2 has been associated with a number of solid cancers, myeloid leukemia, and developmental disorders5,7. However, Shp2 has fallen victim to the same tool shortcomings as described above. To combat these limitations, RapR-Shp2, a specifically and temporally regulatable Shp2 construct, was developed and characterized12.
Prior to the development of RapR-Shp2, it was known that Shp2 was involved in cell migration19,20,21. However, its specific role in the signaling involved in this process was unknown. It was also unknown on what time scale Shp2 regulates the migration of cells and what specific morphodynamic changes it induces at different time points of its activation. Further, it was unclear whether acute and prolonged activation of Shp2 will cause different effects. Using RapR-Shp2, it was found that acute activation of Shp2 induces transient cell spreading, an increase in protrusions, cell polarization, and migration. Specific pathways downstream of Shp2 that regulate distinct morphodynamic changes were also identified12. This protocol provides details for the design and characterization of RapR-Shp2, which can be used to guide the development and application of other RapR phosphatases.
Protocol
1. Design of RapR-phosphatases
- Planning
NOTE: Using RapR-Shp2 as an example, this protocol details the important steps for creating a rapamycin-regulated tyrosine phosphatase. The described protocol is optimized for Shp2 and additional modifications might be needed to fit specific properties of individual phosphatases.- To ensure that the catalytic activity of Shp2 is purely controlled by rapamycin and not by endogenous mechanisms, introduce a mutation into the phosphatase to keep it in a constitutively active conformation. In the case of Shp2, introduce the D61A mutation.
NOTE: If an activating mutation is not introduced for a specific PTPase, then its RapR variant will function as an AND-gated system regulated by endogenous signals and rapamycin. - Identify iFKBP insertion sites in the catalytic domain of the phosphatase using the crystal structure to guide the selection as previously described13. If a structure is not available, perform amino acid sequence alignment with the catalytic domain of Shp2 phosphatase to identify the insertion sites (Figure 2A). Ensure that the insertion site is located in a flexible loop that is structurally coupled to the catalytic pocket but positioned away from the pocket to prevent steric hindrance of substrate binding by iFKBP.
NOTE: Further details of insertion are discussed in the Discussion. - Consider the type of linker sequence to be used between the inserted iFKBP domain and the phosphatase of interest. Choosing the optimal linker sequence is extremely important for the success of the RapR tool and is discussed further in the Discussion (Figure 2B).
- To ensure that the catalytic activity of Shp2 is purely controlled by rapamycin and not by endogenous mechanisms, introduce a mutation into the phosphatase to keep it in a constitutively active conformation. In the case of Shp2, introduce the D61A mutation.
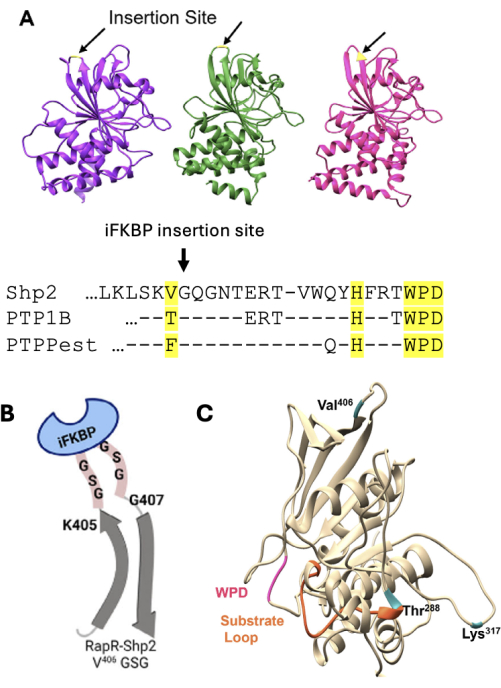
Figure 2: Schematic of considerations when designing RapR-phosphatase. (A) Alignment of Shp2 (purple), PTP1B (green), and PTP-PEST (pink) with the conserved insertion sites highlighted. (B) Representation of linker insertion between Shp2 and iFKBP. (C) Phosphatase domain of Shp2 in beige with insertion sites indicated in blue. This figure was modified from Fauser et al.12. Abbreviations: Shp2 = Src homology-2 domain-containing protein tyrosine phosphatase; iFKBP = insertable FKBP12; PTP = protein tyrosine phosphatase; RapR = Rapamycin-regulated. Please click here to view a larger version of this figure.
2. Creation of the RapR-phosphatase
- Insertion of iFKBP into the phosphatase of interest using modified site-directed mutagenesis
- Generate an iFKBP-encoding "megaprimer" via PCR.
- Design primers for "megaprimer" synthesis that contain 20-25 nucleotides annealing to the 5' or 3' end of iFKPB, a linker sequence that will connect iFKPB to the phosphatase of interest, and 28-32 nucleotides corresponding to the insertion site (Figure 3). Confirm that the resulting product contains the iFKPB flanked by fragments annealing before and after the insertion site in the phosphatase of interest.
- Synthesize the iFKBP containing "megaprimer" via PCR (10 µL of 5x polymerase buffer, 1 µL of 50 ng/µL template DNA containing iFKBP, 2 µL of 10 mM dNTP, 0.5 µL of a 100 µM stock of each "megaprimer" primer [forward and reverse], 35 µL of water, 1 µL DNA polymerase). Split this PCR reaction into 2 separate 25 µL reactions before running the PCR. Run the PCR cycle according to the DNA polymerase manufacturer's recommendations or the following standard PCR cycle: (i) 98 °C for 2 min, (ii) 98 °C for 30 s, (iii) 55-70 °C for 30 s, (iv) 72 °C for 30 s, (v) repeat steps ii-iv 25x, (vi) 72 °C for 2 min.
- Purify the "megaprimer" using agarose gel electrophoresis. Load the PCR contents from the previous step into a 1% agarose gel and apply 120 V for approximately 30 min. Look for the 400 nucleotide-long "megaprimer" in the gel, excise the fragment, and purify using a gel extraction kit (see Table of Materials).
NOTE: The protocol can be paused here.
- Modified site-directed mutagenesis
- Prepare the modified site-directed mutagenesis reaction using a plasmid containing the gene of the phosphatase of interest as the template (5 µL of 5x polymerase buffer, 2 µL of 50 ng/µL template, 2 µL of 10 mM dNTP, 10 µL of extracted "megaprimer", 2.5 µL of water, 2.5 µL of DMSO, 1 µL of DNA polymerase).
NOTE: The addition of DMSO in the reaction mixture lowers the annealing temperature22. - Run the following PCR cycle: initial denaturation step at 98 °C for 3 min, followed by 18 cycles of consecutive steps of incubation at 98 °C for 30 s, 65 °C for 20 s, 60 °C for 20 s, 55 °C for 20 s, 50 °C for 20 s, 72 °C for 18 min. Run the cycle overnight (cycle time is 7 h). Once the cycle is complete, store the PCR reaction at 4 °C.
NOTE: The protocol can be paused here. - After cycle completion, add 1 µL of DpnI enzyme and incubate at 37 °C for 1 h.
NOTE: DpnI will digest the residual template, increasing the yield of the newly synthesized product. - Transform the digested product into DH5α competent cells following the manufacturer’s instructions. Plate the transformation on LB agar plates with the appropriate antibiotic for selection (carbenicillin for the pcDNA RapR-Shp2 construct). Incubate at 37 °C overnight. Once colonies have grown, store the LB plates at 4 °C for up to 1 month.
NOTE: The protocol can be paused here.
- Prepare the modified site-directed mutagenesis reaction using a plasmid containing the gene of the phosphatase of interest as the template (5 µL of 5x polymerase buffer, 2 µL of 50 ng/µL template, 2 µL of 10 mM dNTP, 10 µL of extracted "megaprimer", 2.5 µL of water, 2.5 µL of DMSO, 1 µL of DNA polymerase).
- Generate an iFKBP-encoding "megaprimer" via PCR.
- Isolation of desired DNA construct containing RapR-phosphatase
- Perform a PCR screen of bacterial colonies23. As not all colonies grown on an LB plate will contain the desired plasmid, be sure to screen before the isolation of plasmid DNA.
NOTE: Use primers for the screen that will anneal to the template and in the iFKBP gene. The screen can be done using Taq polymerase mastermix (see Table of Materials). - Once positive colonies are identified, inoculate one positive colony into 3 mL of LB broth including the proper antibiotic, incubate, and shake at 37 °C overnight.
- Extract plasmid DNA from the liquid culture following the manufacturer’s instructions for the plasmid DNA extraction kit (see Table of Materials)
NOTE: It is recommended to sequence the newly created RapR PTPase construct before continuing to in vitro evaluation.
- Perform a PCR screen of bacterial colonies23. As not all colonies grown on an LB plate will contain the desired plasmid, be sure to screen before the isolation of plasmid DNA.
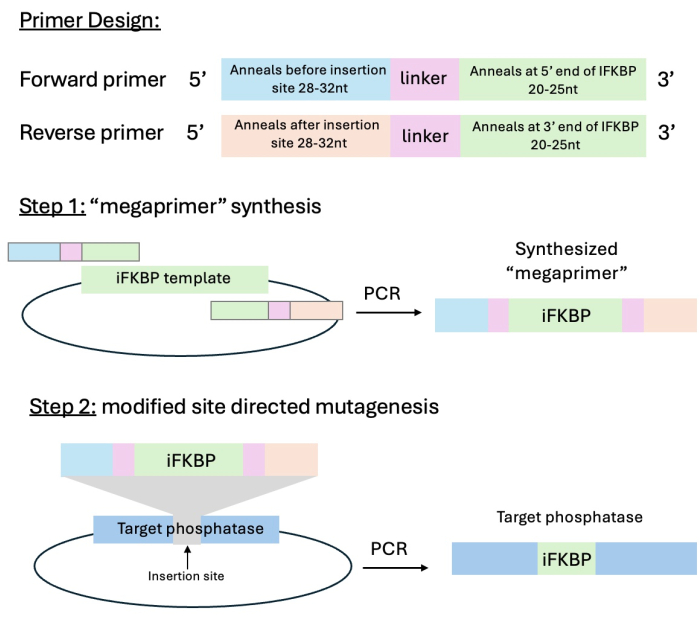
Figure 3: Schematic of primer design and the modified site-directed mutagenesis cloning strategy. Step 1 is the synthesis of the iFKBP containing "megaprimer" with "sticky ends" annealing to the insertion site of the phosphatase of interest, and step 2 is the insertion of the "megaprimer" into the phosphatase of interest. This figure was modified from Karginov et al.13. Abbreviation: iFKBP = insertable FKBP12. Please click here to view a larger version of this figure.
3. Evaluation of RapR-PTPase by in vitro activity assay
NOTE: This protocol is used to assess the regulation of the activity of engineered RapR-PTPase. Below is described the analysis of immunoprecipitated Shp2 using a phosphorylated N-terminal fragment of paxillin as a substrate. A different substrate may need to be selected for a specific PTPase of interest.
- Day 1
- Plate 800,000 HEK293T cells/well in a 6-well tissue culture plate in DMEM media supplemented with L-glutamine supplement (final concentration of 2 mM) and 10% FBS by volume. Place in a 37 °C and 5% CO2 incubator overnight.
- Day 2
- Once the cells are ~60–80% confluent, transfect them according to the preferred transfection regent manufacturer’s protocol (see Table of Materials).
- Example of a transfection protocol: Add 1 µg of plasmid DNA (pcDNA-mCherry-FRB and pcDNA-flag-RapR-Shp2, each) to 200 µL of serum-free DMEM and 6 µL of transfection reagent and incubate the mixture at room temperature for 15 min. Add 100 µL of transfection mixture per well of cells. Include samples transfected with constitutively active and dominant negative Shp2 constructs as controls. Incubate overnight in a 37 °C and 5% CO2 incubator.
- Once the cells are ~60–80% confluent, transfect them according to the preferred transfection regent manufacturer’s protocol (see Table of Materials).
- Day 3
- Preparation of Protein-G Sepharose
NOTE: Prepare cut pipette tips prior to handling Sepharose. Cut tips should be used in every step where Sepharose is handled. Sepharose needs to be thoroughly resuspended in every step prior to pipetting.- Resuspend and take the appropriate amount of Protein-G Sepharose into a 1.5 mL tube. For each well in a 6-well plate, use 10 µL of Protein-G Sepharose. For a full 6-well plate, use 60 µL of Sepharose.
- Wash the Sepharose with 1 mL of Lysis Buffer (see Table of Materials). Microcentrifuge for 1 min at 2,000 × g and carefully remove the Lysis Buffer. Repeat 2x.
- Resuspend the Sepharose in 380 µL of Lysis Buffer. Add BSA to a final concentration 1 mg/mL and antibody (0.5 µL for each IP sample, anti-Flag Antibody [see Table of Materials]). Invert on a rotator at 4 °C for 1.5 h.
NOTE: The optimal amount of antibody should be determined experimentally for each antibody used.
- Prepare cells for immunoprecipitation.
- Add an appropriate amount of rapamycin (1 mM stock solution in ethanol) to a final concentration of 1 µM or add the same volume of ethanol (negative control) to samples. Incubate for 1 h in a 37 °C and 5% CO2 incubator.
- Place the 6-well plate with cells on ice and gently wash the cells with 3 mL of cold PBS. Aspirate the PBS and add 300 µL of Lysis Buffer (with either 1 µM rapamycin for the activated phosphatase sample or an equivalent volume of ethanol in the negative control sample) and scrape with a cell scraper. Transfer the samples to 1.5 mL tubes and microcentrifuge for 10 min at 1,000 × g at 4 °C.
- Transfer 20 µL of the supernatant into a new 1.5 mL tube to check for expression levels. Add 20 µL of 2x Laemmli buffer with 5% β-mercaptoethanol to each tube to be used to check expression and incubate at 95–100 °C for 5 min. Use the remaining supernatant for the immunoprecipitation in step 3.3.3.3.
- Immunoprecipitation of RapR-Shp2
- Wash the Protein-G Sepharose from step 3.3.1.3. with 1 mL of Lysis Buffer. Microcentrifuge for 1 min at 2,000 × g at 4 °C each time and carefully aspirate the Lysis Buffer. Repeat this step 2x.
- Using cut tips, resuspend the Sepharose in Lysis Buffer (50 µL per sample, so for a 6-well plate, resuspend in 300 µL of Lysis Buffer). Pipette 50 µL of the suspended Sepharose mixture into a separate new 1.5 mL tube for each sample.
NOTE: There will be resuspended Sepharose in Lysis Buffer solution left over due to the volume of the Sepharose. - Add the remaining supernatant from the prepared cells from step 3.3.2.3 into each tube of Sepharose. Rotate at 4 °C for 1.5–2 h.
- Phosphatase activity assay
NOTE: The Shp2 substrate should be prepared by phosphorylating purified N-terminal fragment of paxillin using constitutively active Src kinase following the previously described kinase reaction protocol12.- Towards the end of step 3.3.3.3, prepare the phospho-paxillin solution. Per sample, add phospho-paxillin solution (final concentration of 3 µg/mL) to 40 µL of total Imidazole Buffer (see Table of Materials). Add rapamycin to the final concentration of 1 µM to the aliquot of phospho-paxillin solution for activated RapR-Shp2 samples and an equivalent volume of ethanol to the solution for non-activated samples.
- Set the heating shaker to 32 °C.
NOTE: Other phosphatases may need a different temperature for optimal activity. - Wash Sepharose with samples from step 3.3.3.3 2x with 0.5 mL of Lysis Buffer (see Table of Materials). Wash the samples 2x with 0.5 mL of Wash Buffer. Each buffer should have either rapamycin at 1 µM for activated samples or an equivalent volume of ethanol for non-activated samples. Remove as much buffer after the final wash as possible.
NOTE: Once this step is reached, it may be helpful to use a pipette to remove final amounts of buffer slowly and carefully to make sure none of the Sepharose is aspirated. Any leftover buffer will influence the concentration of phospho-paxillin and lead to inaccurate readout. - Add 40 µL of the prepared phospho-paxillin Imidazole Buffer mixture to each sample, with or without rapamycin (sample-dependent). Flick very gently to mix and immediately place in the heating shaker and incubate for 40 min at 32 °C. Set the heating shaker to a minimum of 1,000 rpm to ensure the sample is fully agitated for the duration of the incubation.
- Stop the reaction by adding 40 µL of 2x Laemmli Buffer to each sample. Incubate the samples at 95–100 °C for 5 min. Cool the samples.
- Load 15 µL of each sample (including the lysate samples from step 3.3.2.3.) onto a 4–15% gradient SDS-polyacrylamide gel and run a standard western blot protocol using PVDF membranes.
- Blot using an anti-Flag antibody to detect the RapR-Shp2 construct, anti-mCherry to detect mCherry-FRB, and anti-pY118 or anti-pY31 paxillin to detect paxillin phosphorylation.
NOTE: The resulting western blots should look similar to Figure 4.
- Preparation of Protein-G Sepharose
4. Analysis of RapR-Shp2 activity in living cells
NOTE: This protocol is used to determine the ability of RapR-Shp2 to dephosphorylate endogenous substrates and induce downstream signaling. Other RapR-PTPases will require analysis of their specific substrates and pathways.
- Day 1
- Plate 198,000 A431 cells/well in a 6-well tissue culture plate in DMEM media supplemented with 10% FBS and L-glutamine (final concentration of 2 mM). Place in a 37 °C and 5% CO2 incubator overnight.
- Day 2
- Infect cells with adenoviral constructs expressing RapR-Shp2 and FRB by adding the adenovirus directly to the media already in the dish. Leave the adenovirus on the cells in a 37 °C and 5% CO2 incubator overnight. Use cells infected with FRB only as negative control.
NOTE: The amounts of adenovirus will need to be determined on a case-by-case basis dependent on titer.
- Infect cells with adenoviral constructs expressing RapR-Shp2 and FRB by adding the adenovirus directly to the media already in the dish. Leave the adenovirus on the cells in a 37 °C and 5% CO2 incubator overnight. Use cells infected with FRB only as negative control.
- Day 3
- Starve A431 cells by incubating in DMEM with 0.5% FBS for 4 h before the experiment.
- Add rapamycin to a final concentration of 1 µM or the same volume of ethanol (negative control) to cells for 2–4 h.
NOTE: Incubation could be different for other phosphatases. - Place the 6-well plate with cells on ice and gently wash the cells with 3 mL of cold PBS. Aspirate the PBS, add 300 µL of Lysis Buffer (see Table of Materials), and scrape vigorously with a cell scraper. Transfer the samples to 1.5 mL tubes and microcentrifuge for 5 min at 2,000 × g at 4 °C.
- Transfer 250 µL of each sample to a new 1.5 mL tube. Add 250 µL of 2x Laemmli buffer with 5% β-mercaptoethanol and incubate at 95–100 °C for 5 min.
- Load 25 µL of each sample onto a 4–15% gradient SDS-PAGE gel and run a standard western blot protocol using PVDF membranes.
- Evaluate RapR-Shp2 mediated dephosphorylation of EGFR and PLCγ using anti-pY992 EGFR and anti-pY783 PLCγ antibodies. Assess downstream activation of MAP kinase Erk1/2 by evaluating its phosphorylation on the pT202/pY204 residues.
NOTE: The resulting western blots should look similar to Figure 5.
5. Analyzing morphodynamic changes induced by RapR-Shp2 activation in HeLa cells using live cell imaging
NOTE: This protocol is used to determine the effect of RapR-Shp2 activation on the formation of cell protrusions, cell spreading, and migration.
- Day 1
- Plate 700,000 HeLa cells in a 35 mm cell culture dish and place in a 37 °C and 5% CO2 incubator. Allow the cells to adhere to the dish (~2–3 h).
- Transfect the cells according to the preferred transfection reagent manufacturer’s protocol (see Table of Materials).
- Example transfection protocol: Add 0.5 µg of DNA (pcDNA-mCherry-FRB and pcDNA-Venus-RapR-Shp2 plasmids, each) to 100 µL of serum-free DMEM and 6 µL of transfection reagent. Incubate at room temperature for 15 min. Add the transfection mixture to the cells and incubate overnight in a 37 °C and 5% CO2 incubator. Use cells transfected with pcDNA-mCherry-FRB only as a negative control.
- Coat a live imaging #1.5 25 mm round glass coverslip with fibronectin (5 mg/L) by incubating for 1 h at 37 °C.
- Day 2
- Wash the coated glass coverslip with PBS.
- Plate the transfected HeLa cells on the coated glass coverslip in DMEM media supplemented with 10% FBS and L-glutamine (final concentration of 2 mM) to achieve 30% confluency 4 h prior to imaging.
- Two hours after plating, gently switch the media in the plate to prewarmed serum-free Leibovitz L-15 media for 2 h.
- Stain the cells with CellMask Deep Red plasma membrane stain according to the manufacturer’s protocol.
- Place the coverslip into a clean imaging cell chamber (see Table of Materials). Add 1 mL of prewarmed L-15 media and cover with 1 mL of prewarmed mineral oil to prevent evaporation. Keep the chamber at 37 °C until imaging.
- Image the cells at 37 °C using a heated stage.
- Select proper channels for imaging of the RapR construct and CellMask for epifluorescence imaging. To follow this protocol, use the referenced 40x objective (see Table of Materials) and the following filter sets: 514/10 excitation and 540/21 emission filters for Venus-RapR-Shp2 imaging, 561/10 excitation and 595/40 emission filters for mCherry-FRB imaging, and 640/20 excitation and 655LP emission filters for CellMask Deep Red imaging. Use a multiband polychroic mirror for all fluorescent channels (See Table of Materials).
- Capture images every 2 min for 4–4.5 h.
NOTE: Faster morphodynamic changes will require more frequent acquisition. - After 1 hour of imaging, add rapamycin to a final concentration of 1 µM to stimulate RapR-Shp2.
6. Image analysis
NOTE: This protocol will describe the creation of cell masks based on .TIF stack files collected from live imaging experiments. It will then describe how to create a Macro in ImageJ to analyze the masks, which will result in a spreadsheet of cell area that is then analyzed for changes over time. Finally, cell protrusive and retractive activity will be analyzed using Metamorph.
- Generate a binary mask of CellMask images by using the MovThresh function from CellGeo software package24.
- Copy the MovThresh folder into the image analysis folder.
- Copy the CellMask or other membrane marker channel images into the MovThresh folder.
- If multiple cells were imaged at a single position, crop the images to generate files containing only a single cell prior to creating the mask in MatLab.
- Ensure the directory is set correctly to the MovThresh folder in MatLab.
- Run MovThresh.
- Import image stacks one at a time.
- Select Smoothed Curve and then run Rethreshold.
- Save as Mask_Image Number in a new folder labeled Masks.
- Once all image stacks have been converted into Masks, open the image stacks in ImageJ software25.
- Cell area analysis in ImageJ
- Open all Mask image stacks in ImageJ.
- Adjust Set Measurements to Area.
- Click Plugins | Macros | Record.
- Click Process | Binary | Make Binary.
- Select the default option.
NOTE: Do not click anything extra while recording the Macro. If additional steps are included, it is recommended to restart from step 6.9.1.3.
- Select the default option.
- Click Analyze | Analyze Particles.
- Uncheck all boxes but Summary.
- Finish recording and click Create the make the Macro.
- Hit Run for each image stack after selecting it and save the results table for each image stack. The default option will save as a csv file.
- Save the Macro and re-use for future analyses (optional).
- Processing area data in a spreadsheet
- For each cell analyzed, average the area of the cell from images taken before activation with rapamycin.
- Divide all area values by the average area of the cell preceding rapamycin addition.
- Calculate the average value for each time point for all imaged cells and determine the 90% confidence intervals. Plot the data.
- Cell protrusive activity analysis
- Copy the ProActive folder into the image analysis folder.
- Copy the Masked image stacks from MovThresh into the newly created ProActive folder.
- Open MatLab and set the directory to the ProActive folder.
- Run ProActive.
- Import the Masked cell image stack files into the ProActive GUI.
- Define the type of activity. Protrusive activity analysis will generate area gained data; retractive activity analysis will generate area lost data; and total activity will generate overall area change data (both lost and gained).
- Set area to Define Normalization.
- Smooth curve either using the suggested threshold values or using mean to rethreshold each time point. This averages the thresholds over a specific window, minimizing inconsistencies in thresholding between time points.
- Click File | Save As | Results (.mat)
- If processing all three types of area activity, name the processed data files as follows: ProResultData #, RetResultData #, or ResultData #.
NOTE: Include the space before the number and number the data starting at 1.
- If processing all three types of area activity, name the processed data files as follows: ProResultData #, RetResultData #, or ResultData #.
- Repeat from step 6.9.3.6. for all masked image stacks imported.
- Once all the cells have been processed, close the ProActive GUI.
- Open DataToFileTotalArea in the Matlab editor to process total activity “ResultData” files.
- Change the number of cells to match how many result files have been generated. If 7 cells were processed in steps 6.9.3.5–6.9.3.10 ensure that the first three lines of the script are as follows:
filename=’Result Data ‘;
cells=1:7;
lag=1; - Hit Run and repeat this for DataToFileProtArea and DataToFileRetArea to process protrusive and retractive data, respectively. This analysis will produce three text files “AllCells.txt”, “AllCellsPro.txt”, and “AllCellsRet.txt”
- Open the .txt files in a spreadsheet and normalize as in section 6.9.2. Calculate the average value for each time point for all imaged cells and determine 90% confidence intervals. Plot the data.
NOTE: The resulting plots from the above steps should look similar to Figure 6.
- Cell area analysis in ImageJ
Results
Figure 4 demonstrates results that can be expected from the paxillin-based phosphatase activity assay. In this experiment, constitutively active and dominant negative Shp2 phosphatase activity was compared to that of active and inactive RapR-Shp2 using phospho-paxillin as the readout. The Shp2 constructs were immunoprecipitated and subjected to the activity assay as described in the protocol. The phospho-paxillin readouts for constitutively active Shp2 and active RapR-Shp2 were similar,...
Discussion
This protocol provides detailed steps for the development, characterization, and application of chemogenetically controlled phosphatases. The RapR-Shp2 tool relies on a rapamycin-regulated switch inserted in the Shp2 catalytic domain. The strength of this tool is the specificity and tight temporal control of phosphatase activity. The tool is applicable to other phosphatases and, in combination with previously described RapR-TAP technology, allows for the reconstruction of individual downstream signaling pathways
Disclosures
The authors have no conflicts of interest to disclose.
Acknowledgements
The authors acknowledge Dr. Jordan Fauser for her contribution to the development of RapR-Shp2 and associated protocols. The work was supported by a 5R35GM145318 award from NIGMS, an R33CA258012 award from NCI, and a P01HL151327 award from NHLBI.
Materials
| Name | Company | Catalog Number | Comments |
| #1.5 Glass Coverslips 25 mm Round | Warner Instruments | 64-0715 | |
| 1.5 mL Tubes | USA Scientific | cc7682-3394 | |
| 2x Laemmli Buffer | For 500 mL: 5.18 g Tris-HCL, 131.5 mL glycerol, 52.5 mL 20% SDS, 0.5 g bromophenol blue, final pH 6.8 | ||
| 4-20% Mini-PROTEAN TGX Precast Gel | Biorad | 4561096 | |
| 5x Phusion Plus Buffer | Thermo Scientific | F538L | |
| A431 Cells | ATCC | CRL-1555 | |
| Agarsose | GoldBiotech | A-201 | |
| Attofluor Cell Chamber | invitrogen | A7816 | |
| Benchmark Fetal Bovine Serum (FBS) | Gemini Bio-products | 100-106 | Heat Inactivated Triple 0.1 µm sterile-filtered |
| Brig 35,30 w/v % | Acros | 329581000 | |
| BSA | GoldBiotech | A-420 | |
| CellGeo | N/A | N/A | Published in 10.1083/jcb.201306067 |
| CellMask Deep Red plasma membrane dye | invitrogen | c10046 | |
| Colony Screen MasterMix | Genesee | 42-138 | |
| DH5a competent cells | NEB | C2987H | |
| DMEM | Corning | 15-013-CV | |
| DMSO | Thermo Scientific | F-515 | |
| DNA Ladder | GoldBio | D010-500 | |
| dNTPs | NEB | N04475 | |
| DpnI Enzyme | NEB | R01765 | |
| DTT | GoldBio | DTT10 | DL-Dithiothreitol, Cleland's Reagents |
| EGTA | Acros | 409910250 | |
| Fibronectin from bovine plasma | Sigma | F1141 | |
| FuGENE(R) 6 Transfection Reagent | Promega | E2692 | transfection reagent |
| Gel extraction Kit | Thermo Scientific | K0692 | GeneJET Gel Extraction Kit |
| Gel Green Nucleic Acid Stain | GoldBio | G-740-500 | |
| Gel Loading Dye Purple 6x | NEB | B7024A | |
| Glutamax | Gibco | 35050-061 | GlutaMAX-l (100x) 100 mL |
| HEK 293T Cells | ATCC | CRL-11268 | |
| HeLa Cells | ATCC | CRM-CCL-2 | |
| HEPES | Fischer | BP310-500 | |
| ImageJ Processing Software | N/A | N/A | |
| Igepal CA-630 (NP40) | Sigma | I3021 | |
| Imidazole Buffer | 25 mM Imidazole pH 7.2, 2.5 mM EDTA, 50 mM NaCl, 5 mM DTT | ||
| KCl | Sigma | P-4504 | |
| L-15 1x | Corning | 10-045-CV | |
| LB Agar | Fisher | BP1425-2 | |
| Lysis Buffer | 20 mM Hepes-KOH, pH 7.8, 50 mM KCl, 1 mM EGTA, 1% NP40 | ||
| MATLAB | MathWorks | N/A | R 2022b update was used to run CellGeo functions |
| Metamorph Microscopy Automation and Image Analysis Software | N/A | N/A | |
| MgCl2 | Fisher Chemical | M33-500 | |
| Mineral Oil | Sigma | M5310 | |
| MiniPrep Kit | Gene Choice | 96-308 | |
| Mini-PROTEAN TGX Precast Gels 12 well | Bio-Rad | 4561085 | |
| Molecular Biology Grade Water | Corning | 46-000-CV | |
| Multiband Polychroic Mirror | Chroma Technology | 89903BS | |
| NaCl | Fisher Chemical | S271-3 | |
| Olympus UPlanSAPO 40x objective | Olympus | N/A | |
| PBS w/o Ca and Mg | Corning | 21-031-CV | |
| PCR Tubes | labForce | 1149Z65 | 0.2 mL 8-Strip Tubes and Caps, Rigid Strip Individually Attached Dome Caps |
| Phusion Plus DNA Polymerase | Thermo Scientific | F630S | |
| Primers | IDT | ||
| Protein-G Sepharose | Millipore | 16-266 | |
| PVDF Membranes | BioRad | 1620219 | Immun-Blot PVDF/Filter Paper Sandwiches |
| Rapamycin | Fisher | AAJ62473MF | |
| 0.25% Trypsin, 2.21 mM EDTA, 1x [-] sodium | Corning | 25-053-CI | |
| Tris-Acetate-EDTA (TAE) 50x | Fischer | BP1332-1 | for electrophoresis |
| Wash Buffer | 20 mM Hepes-KOH, pH 7.8, 50 mM KCl, 100 mM NaCl, 1 mM EGTA, 1% NP40 | ||
| β-Mercaptoethanol | Fisher Chemical | O3446I-100 | |
| Antibodies | |||
| Anti-EGFR Antibody | Cell Signaling | 2232 | |
| Anti-Erk 1/2 Antibody | Cell Signaling | 9102 | |
| Anti-Flag Antibody | Millipore-Sigma | F3165 | |
| Anti-GAPDH Antibody | invitrogen | AM4300 | |
| Anti-GFP Antibody | Clontech | 632380 | |
| Anti-mCherry Antibody | invitrogen | M11217 | |
| Anti-paxillin Antibody | Thermo Fischer | BDB612405 | |
| Anti-phospho-EGFR Y992 Antibody | Cell Signaling | 2235 | |
| Anti-phospho-Erk 1/2 T202/Y204 Antibody | Cell Signaling | 9101 | |
| Anti-phospho-paxillin Y31 Antibody | Millipore-Sigma | 05-1143 | |
| Anti-phospho-PLCγ Y783 Antibody | Cell Signaling | 14008 | |
| Anti-PLCγ Antibody | Cell Signaling | 5690 |
References
- Amin, A. R. M. R., et al. SHP-2 tyrosine phosphatase inhibits p73-dependent apoptosis and expression of a subset of p53 target genes induced by EGCG. Proc Natl Acad Sci U S A. 104 (13), 5419-5424 (2007).
- Niogret, C., et al. Shp-2 is critical for ERK and metabolic engagement downstream of IL-15 receptor in NK cells. Nat Commun. 10, 1-14 (2019).
- Wang, Y. -. C., et al. Helicobacter pylori infection activates Src homology-2 domain-containing phosphatase 2 to suppress IFN-γ signaling. J Immunol. 193 (8), 4149-4158 (2014).
- Yu, M., et al. The tyrosine phosphatase SHP2 promotes proliferation and oxaliplatin resistance of colon cancer cells through AKT and ERK. Biochem Biophys Res Commun. 563, 1-7 (2021).
- Lauriol, J., et al. Developmental SHP2 dysfunction underlies cardiac hypertrophy in Noonan syndrome with multiple lentigines. J Clin Invest. 126 (8), 2989-3005 (2016).
- Tartaglia, M., et al. Mutations in PTPN11, encoding the protein tyrosine phosphatase SHP-2, cause Noonan syndrome. Nat Genet. 29, 465-468 (2001).
- Xu, R., et al. Overexpression of Shp2 tyrosine phosphatase is implicated in leukemogenesis in adult human leukemia. Blood. 106 (9), 3142-3149 (2005).
- Mustelin, T., et al. Protein tyrosine phosphatases. Front Biosci. 7 (4), d85-d142 (2002).
- Stanford, S. M., Bottini, N. Targeting tyrosine phosphatases: time to end the stigma. Trends Pharmacol Sci. 38 (6), 524-540 (2017).
- Guo, M., et al. Targeting phosphatases: From molecule design to clinical trials. Eur J Med Chem. 264, 116031 (2024).
- Weiser, D. C., Shenolikar, S. Use of protein phosphatase inhibitors. Curr Protoc Mol Biol. 62, 18.10.1-18.10.13 (2003).
- Fauser, J., et al. Dissecting protein tyrosine phosphatase signaling by engineered chemogenetic control of its activity. J Cell Biol. 221 (8), e202111066 (2022).
- Karginov, A. V., Hahn, K. M. Allosteric activation of kinases: design and application of RapR kinases: design and application of RapR kinases. Curr Protoc Cell Biol. Chapter 14 (Unit 14.13), (2011).
- Karginov, A. V., Ding, F., Kota, P., Dokholyan, N. V., Hahn, K. M. Engineered allosteric activation of kinases in living cells. Nat Biotechnol. 28, 743-747 (2010).
- Dagliyan, O., Dokholyan, N. V., Hahn, K. M. Engineering proteins for allosteric control by light or ligands. Nat Protoc. 14, 1863-1883 (2019).
- Wang, Y., et al. Targeting the SHP2 phosphatase promotes vascular damage and inhibition of tumor growth. EMBO Mol Med. 13, e14089 (2021).
- Wang, L., et al. SHP2 regulates the development of intestinal epithelium by modifying OSTERIX+ crypt stem cell self-renewal and proliferation. FASEB J. 35, e21106 (2021).
- Zhang, E. E., Chapeau, E., Hagihara, K., Feng, G. -. S. Neuronal Shp2 tyrosine phosphatase controls energy balance and metabolism. Proc Natl Acad Sci U S A. 101 (45), 16064-16069 (2004).
- Song, Y., Zhao, M., Zhang, H., Yu, B. Double-edged roles of protein tyrosine phosphatase SHP2 in cancer and its inhibitors in clinical trials. Pharmacol Ther. 230, 107966 (2022).
- Hartman, Z. R., Schaller, M. D., Agazie, Y. M. The tyrosine phosphatase SHP2 regulates focal adhesion kinase to promote EGF-induced lamellipodia persistence and cell migration. Mol Cancer Res. 11 (6), 651-664 (2013).
- Yu, D. H., Qu, C. K., Henegariu, O., Lu, X., Feng, G. S. Protein-tyrosine phosphatase Shp-2 regulates cell spreading, migration, and focal adhesion. J Biol Chem. 273 (33), 21125-21131 (1998).
- Kramer, M. F., Coen, D. M. Enzymatic amplification of DNA by PCR: standard procedures and optimization. Curr Protoc Cell Biol. 10, A.3F.1-A.3F.14 (2001).
- Sandhu, G. S., Precup, J. W., Kline, B. C. Rapid one-step characterization of recombinant vectors by direct analysis of transformed Escherichia coli colonies. Biotechniques. 7, 689-690 (1989).
- Tsygankov, D., et al. CellGeo: A computational platform for the analysis of shape changes in cells with complex geometries. J Cell Biol. 204 (3), 443-460 (2014).
- Schneider, C. A., Rasband, W. S., Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 9, 671-675 (2012).
- Ray, A. -. M., Klomp, J. E., Collins, K. B., Karginov, A. V., Tan, A. -. C., Huang, P. H. Dissecting kinase effector signaling using the RapRTAP methodology. Kinase Signaling Networks. , 21-33 (2017).
- Mercan, F., Bennett, A. M. Analysis of protein tyrosine phosphatases and substrates. Curr Protoc Mol Biol. 18 (Unit-18.16), (2010).
- Karginov, A. V., et al. Light regulation of protein dimerization and kinase activity in living cells using photocaged rapamycin and engineered FKBP. J Am Chem Soc. 133 (3), 420-423 (2011).
- Karginov, A. V., Hahn, K. M., Deiters, A. Optochemical activation of kinase function in live cells. Methods Mol Biol. 1148, 31-43 (2014).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved