A subscription to JoVE is required to view this content. Sign in or start your free trial.
Determining the Serum Stability of Human Adenosine Deaminase 1 Enzyme
In This Article
Summary
In this article, we detail methods to characterize an enzyme's ability to retain function when incubated at 37 °C in human serum, a pharmacological property referred to as its serum stability. This ability may be a key factor in predicting an enzyme's pharmacokinetic profile and its suitability for therapeutic use.
Abstract
The concept of enzyme stability is typically used to refer to an enzyme's thermostability - its ability to retain structure and activity as temperature increases. For a therapeutic enzyme, other measures of stability may also be critical, particularly its ability to retain function in human serum at 37 °C, which we refer to as serum stability. Here, we describe an in vitro assay to assess the serum stability of the wildtype Homo sapiens adenosine deaminase I (HsADA1) enzyme using an absorbance-based microplate procedure. Specifically, this manuscript describes the preparation of buffers and reagents, a method arranging for the coincubation of HsADA1 in serum, a method to analyze the test samples using a microplate reader, and an accompanying analysis to determine the fraction of activity that an HsADA1 enzyme retains in serum as a function of time. We further discuss considerations to adapt this protocol to other enzymes, using an example of a Homo sapiens kynureninase enzyme, to help aid the protocol's adaptation to other enzymes where serum stability is of interest.
Introduction
The following method allows a user to quantitatively assess an enzyme's ability to retain its activity when exposed to conditions that mimic what it will encounter following intravenous injection. The in vitro method mimics such in vivo conditions and consists of the incubation of the enzyme in pooled human serum at 37 °C and time-coursed analyses of retention of enzyme activity. We refer to an enzyme's ability to retain activity in these conditions as its serum stability, and the analysis method for enzyme activity takes advantage of differences in absorbance between an enzyme's substrate and the resulting product. The concept of serum stability is not just enzyme-specific and has been applied to several other treatment modalities as well. For example, the serum stability of RNA aptamers targeting COVID spike proteins has previously been assessed by monitoring their degradation post-incubation with fetal calf serum1. Antibacterial peptides have also been assessed for their ability to suppress bacterial growth post-incubation with pooled human serum2.
HsADA1 is an enzyme that catalyzes the conversion of adenosine or 2-deoxyadenosine into inosine or 2-deoxyinosine, respectively3. Adenosine has an absorption peak of 260 nm, while inosine absorbs strongest at 250 nm4. This shift in absorption peaks can be detected on a microplate reader by a decrease in absorption intensity at 260 nm when HsADA1 is added to adenosine. The HsADA1 enzyme has important implications in the human body, and its deficiency causes a severe combined immunodeficiency (ADA-SCID)5. The bovine homolog of HsADA1, BtADA1, can be utilized as an enzyme replacement therapeutic for the treatment of ADA-SCID, and we have previously shown that wildtype HsADA1 loses its activity when incubated with pooled human serum, potentially hindering its use as a therapeutic6. Therefore, we selected the wildtype HsADA1 enzyme to demonstrate a procedure to determine an enzyme's serum stability. A detailed purification method for HsADA1 has been described previously6.
In the following protocol (as detailed in Figure 1), we demonstrate how to co-incubate wildtype HsADA1 in pooled human serum at 37 °C. During this time, test samples are taken at defined time points and flash-frozen for future analysis. Once all samples have been taken, a microplate assay is run whereby each sample is combined with the substrate, with the resulting absorbance decrease being a correlation for the retained activity of the enzyme. Representative results illustrating the serum stability of HsADA1 are shown, and because this metric may be relevant to determining the potential therapeutic value of other enzymes, we also discuss considerations to adapt this protocol to an engineered Homo sapiens kynureninase enzyme (HsKYNase) and any enzymes more generally. HsKYNase is an enzyme involved in the metabolism of tryptophan and is able to degrade the tryptophan-byproducts kynurenine and hydroxy-kynurenine (OH-Kyn) into anthranilic acid and hydroxy-anthranilic acid (OH-AA), respectively. Enzyme-mediated modulation of tryptophan metabolism may be of therapeutic relevance7.
Protocol
1. Serum incubation
- Prepare a 10x stock of HsADA1 in 1x PBS pH 7.4 (1x PBS) at a final concentration of 10 µM. Thaw a 15 mL aliquot of pooled human serum and pre-warm it to 37 °C. Prewarm a 50 mL aliquot of 1x PBS to 37 °C.
- Prepare the enzyme-serum incubation mixtures by adding 100 µL of the 10x HsADA1 stock to 900 µL of pooled human serum in a low-bind microcentrifuge tube, referred to as the enzyme + serum mixture. In a separate low-bind microcentrifuge tube, add 100 µL of the 10x HsADA1 stock to 900 µL of 1x PBS for comparison to a non-serum control, referred to as the enzyme + 1x PBS mixture. The final concentration of HsADA1 in each mixture will be 1 µM.
- Prepare an additional control in a low-bind microcentrifuge tube consisting of 100 µL of 1x PBS mixed with 900 µL of pooled human serum. This will serve as the non-enzyme control for the enzyme + serum mixture. Additionally, make a control consisting of 1 mL of 1x PBS. This will serve as the non-enzyme control for the enzyme + 1x PBS mixture.
- Aliquot 100 µL of each mixture from step 1.2 into separate low-bind tubes and dilute 1:1 with 100 µL of 30% (v/v) glycerol in 1x PBS. Flash freeze with liquid nitrogen and store at -80 °C. These samples will be the Day 0 timepoint, and the concentration of HsADA1 in them will be 500 nM. Repeat this step for the non-enzyme control mixtures from 1.3. Seal all enzymes and control mixtures with transparent film and place them into a 37 °C incubator.
- After 24 h, aliquot 100 µL of each incubated sample and dilute 1:1 with 100 µL of 30% (v/v) glycerol in 1x PBS. Flash freeze with liquid nitrogen and store at -80 °C. These samples will be the Day 1 timepoint. Return all enzyme and control mixtures to the 37 °C incubator. Repeat this step at 72 h and 120 h.
NOTE: Aliquoted samples do not necessarily need to be flash-frozen. Alternatively, the user may perform the microplate-based assay immediately after taking each sample, although fresh standard curves and adenosine substrate stocks should be prepared each time a sample is tested.
2. Microplate-based assay
- Prepare the microplate reader by setting up a kinetic read protocol (Figure 2) to measure the absorbance at 260 nm with the shortest reading interval for 30 min and preheat the reader to a setpoint of 37 °C.
- Thaw all collected samples on ice. Dilute each thawed sample 1:10 using 1x PBS prewarmed to 37 °C as a diluent in low-bind microcentrifuge tubes. The new HsADA1 concentration in these samples will be 50 nM.
- Make a 10x dilution into 1x PBS from the non-enzyme serum control sample as well. Additionally, prepare a 5 mM adenosine stock by adding 66.8 mg of adenosine to 50 mL of 1x PBS. Prepare a 250 µM dilution of adenosine by adding 400 µL of the adenosine stock to 7,600 µL of 1x PBS and prewarm it to 37 °C in an incubator.
- To a 96-well UV-compatible microplate, add 160 µL of the 250 µM adenosine dilution and 40 µL of each diluted protein sample in triplicate for a total volume of 200 µL. This will yield a final adenosine concentration of 200 µM and a final enzyme concentration of 10 nM.
- Repeat the dilution for the control samples by adding 40 µL of the diluted serum or 1x PBS controls and 160 µL of the 250 µM adenosine dilution in triplicate. This will also yield a final adenosine concentration of 200 µM but without enzyme.
NOTE: It may be beneficial to add all diluted enzyme and control samples individually to the microplate first and then add the adenosine substrate with a 12-channel micropipette to reduce the total loading time. Additionally, UV-transparent plates must be used in this step to prevent the plate itself from confounding the measured absorbance signal. - Measure the absorbance of each well at 260 nm for 30 min at 37 °C. Once done, export the data to a spreadsheet for further analysis.
3. Analysis of microplate reader data
- Determine the slope of the linear region of the absorbance data as a function of time for the first several minutes of the microplate data. For the serum-diluted samples, this corresponded to the first ~120 s. For the 1x PBS-diluted samples, this corresponded to the first ~270 s. This slope of declining absorbance values will have a negative numerical value and will have units of change in absorbance units per second (ΔA260/s).
- Subtract out the slope of the negative control consisting of pooled human serum from the slope of the enzyme + serum data. Perform the same operation on the enzyme + 1x PBS mixtures by subtracting out the slope of the 1x PBS control.
- Normalize all the adjusted slopes to the original slope at t=0 h to obtain the fraction of activity remaining using Equation 1.
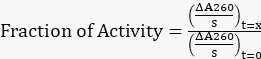 Equation 1
Equation 1 - Plot the fraction of activity remaining versus time.
Results
The figures show the results of the assay run when conducted with wildtype HsADA1. Figure 3A,B illustrate the absorbance decline curves at 260 nm of the samples originating from the 1x PBS/serum-enzyme mixtures for wildtype HsADA1 after the addition of adenosine. This declining absorbance as a function of time data is what the user may expect upon successful completion of the microplate-based assay and is similar to absorbance data that would arise after adding sufficient am...
Discussion
This protocol uses absorbance change as the substrate is converted to the product to gauge the activity of an enzyme. As such, the substrate and product must have distinct spectral profiles. This is the case with adenosine and inosine both having distinct spectral profiles and extinction coefficients between 260-265 nm6,8,12,13. This assay is inspired by several previous works. Kalackar, for ex...
Disclosures
JB and MRJ are inventors on patents or patent applications related to adenosine deaminase and/or kynureninase enzymes. All other authors have no conflicts of interest to disclose.
Acknowledgements
This work was supported by the National Institutes of Health [1DP2CA280622-01] and funding from Biolocity. We thank Dr. Maria Jennings and Andrea Fox for providing the HsADA1 and HsKYNase expression vectors.
Materials
| Name | Company | Catalog Number | Comments |
| Adenosine | Sigma Aldrich | A9251-25G | 25 g |
| BioTek Synergy HT Microplate Reader | |||
| Eppendorf LoBind Microcentrifuge Tubes: Protein | Fisher Scientific | 13-698-795 | 2 mL |
| Glycerol | Fisher Scientific | G33-4 | 4 L |
| HsKYNase66-W102H-T333N | In-house | ||
| Human Serum, Pooled | MP Biomedicals | 92931149 | 100 mL |
| Hydroxy-kynurenine | Cayman Chemicals | 27778 | |
| Inosine | TCI | I0037 | 25 g |
| PBS, 1x pH 7.4+/- 0.1 | Corning | 21-040-CM | |
| Pyridoxal 5-phosphate monohydrate, 99% | Thermo Scientifc | 228170010 | 1 g |
| UV-STAR MICROPLATE, 96 WELL, COC, F-BOTTOM | Greiner Bio | 655801 | |
| Wildtype Human Adenosine Deaminase 1 | In-house |
References
- Valero, J., et al. A serum-stable RNA aptamer specific for SARS-COV-2 neutralizes viral entry. Proc Natl Acad Sci U S A. 118 (50), e2112942118 (2021).
- Iannuzo, N., et al. High-throughput screening identifies synthetic peptides with antibacterial activity against mycobacterium abscessus and serum stability. ACS Omega. 7 (27), 23967-23977 (2022).
- Ma, M. T., Jennings, M. R., Blazeck, J., Lieberman, R. L. Catalytically active holo homo sapiens adenosine deaminase i adopts a closed conformation. Acta Crystallograph Sect D. 78 (1), 91-103 (2022).
- Li, W., et al. Determination of 4 nucleosides via one reference compound in chinese cordyceps by hplc-uv at equal absorption wavelength. Natural Prod Comm. 18 (3), 1934578X231161410 (2023).
- Whitmore, K. V., Gaspar, H. B. Adenosine deaminase deficiency - more than just an immunodeficiency. Front Immunol. 7, 314 (2016).
- Jennings, M. R., et al. Optimized expression and purification of a human adenosine deaminase in E. coli and characterization of its asp8asn variant. Prot Express Purificat. 213, 106362 (2024).
- Blazeck, J., et al. Bypassing evolutionary dead ends and switching the rate-limiting step of a human immunotherapeutic enzyme. Nat Catalysis. 5 (10), 952-967 (2022).
- Lu, J., Grenache, D. G. Development of a rapid, microplate-based kinetic assay for measuring adenosine deaminase activity in body fluids. Clinica Chimica Acta. 413 (19), 1637-1640 (2012).
- Maciel, L. G., Dos Anjos, J. V., Soares, T. A. Fast and low-cost evaluation of hydroxykynurenine activity. MethodsX. 7, 100982 (2020).
- Bokman, A. H., Schweigert, B. S. 3-hydroxyanthranilic acid metabolism. Iv. Spectrophotometric evidence for the formation of an intermediate. Arch Biochem Biophys. 33 (2), 270-276 (1951).
- Tanford, C. . Advances in protein chemistry. 23, 121-282 (1968).
- Tritsch, G. L. Validity of the continuous spectrophotometric assay of kalckar for adenosine deaminase activity. Anal Biochem. 129 (1), 207-209 (1983).
- Gracia, E., et al. The catalytic site structural gate of adenosine deaminase allosterically modulates ligand binding to adenosine receptors. FASEB J. 27 (3), 1048-1061 (2013).
- Kalckar, H. M. Differential spectrophotometry of purine compounds by means of specific enzymes: Iii. Studies of the enzymes of purine metabolism. J Bio Chem. 167 (2), 461-475 (1947).
- Hartwick, R., Jeffries, A., Krstulovic, A., Brown, P. R. An optimized assay for adenosine deaminase using reverse phase high pressure liquid chromatography. J Chromatographic Sci. 16 (9), 427-435 (1978).
- Paul, M. K., Grover, V., Mukhopadhyay, A. K. Merits of hplc-based method over spectrophotometric method for assessing the kinetics and inhibition of mammalian adenosine deaminase. J Chromatography B. 822 (1), 146-153 (2005).
- Ubbink, J. B., Vermaak, W. J. H., Bissbort, S. H. High-performance liquid chromatographic assay of human lymphocyte kynureninase activity levels. J Chromatography B: Biomed Sci Appl. 566 (2), 369-375 (1991).
- Tsentalovich, Y. P., Snytnikova, O. A., Forbes, M. D. E., Chernyak, E. I., Morozov, S. V. Photochemical and thermal reactivity of kynurenine. Exp Eye Res. 83 (6), 1439-1445 (2006).
- Demain, A. L., Vaishnav, P. Production of recombinant proteins by microbes and higher organisms. Biotechnol Adv. 27 (3), 297-306 (2009).
- Gräslund, S., et al. Protein production and purification. Nat Methods. 5 (2), 135-146 (2008).
- Faber, M. S., Whitehead, T. A. Data-driven engineering of protein therapeutics. Curr Opin Biotechnol. 60, 104-110 (2019).
- Das, S., Zhao, L., Elofson, K., Finn, M. G. Enzyme stabilization by virus-like particles. Biochemistry. 59 (31), 2870-2881 (2020).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved