A subscription to JoVE is required to view this content. Sign in or start your free trial.
Measuring Enzymatic Activity of Neurodevelopmental Disorder-Associated Deubiquitylating Enzymes via an In Vitro Ubiquitin Chain Cleavage Assay
In This Article
Summary
This protocol explains how to measure the effect of genetic variation associated with neurodevelopmental disorders on deubiquitylating enzyme activity by combining recombinant protein purification with ubiquitin chain cleavage assays.
Abstract
Neurodevelopmental disorders (NDDs) are associated with impairments in nervous system function but often remain poorly understood at the molecular level. Discrete disorders caused by single genes provide models to investigate mechanisms driving atypical neurodevelopment. Variants of genes encoding deubiquitylating enzyme (DUB) family proteins are associated with several NDDs, but there is a need to determine the pathogenic mechanisms of disorders driven by these gene variants. The impact of gene variants on DUB activity can be experimentally determined using a substrate-independent in vitro ubiquitin cleavage assay. This assay does not require knowledge of downstream substrates to directly measure catalytic activity. Here, the protocol for determining the impact of gene variants on enzymatic activity is modeled using the DUB Ubiquitin Specific Protease 27, X-linked (USP27X), which is mutated in X-linked intellectual disability 105 (XLID105). This experimental pipeline can be used to clarify the mechanisms underlying neurodevelopmental disorders driven by variants in DUB genes.
Introduction
Neurodevelopmental disorders (NDDs) arise from diverse etiologies with environmental or genetic determinants that drive atypical nervous system development1. Next-generation sequencing genetic testing has linked an increasing number of variants in ubiquitin system-related genes with genetic NDDs2. The ubiquitin system catalyzes the ligation of the small protein modifier ubiquitin to primarily lysine residues in protein substrates to drive changes in cellular behavior, including localization, stability, protein-protein interactions, or activity3. Ubiquitylation is mediated by E1 activating, E2 conjugating, and E3 ligase enzymes4 and is reversible by the activity of deubiquitylating enzymes (DUBs) that catalyze the cleavage and removal of ubiquitin from protein substrates5. Ubiquitin can be ligated to the substrate as a monomer (monoubiquitylation) or polymeric chains (polyubiquitylation) that are formed on any of the seven lysine residues (K6, K11, K27, K29, K33, K48, K63) or the M1 residue of ubiquitin. These different ubiquitin chain topologies and their combinations create a cellular code that is key for signal transduction6.
DUBs such as USP27X, USP7, USP9X, USP48, STAMBP, OTUD4, OTUD6B, and OTUD5 have been associated with NDDs2,7,8,9,10,11. For most NDDs, the molecular mechanisms that drive pathogenesis remain experimentally undefined. Some of the DUBs driving recently described disorders are poorly understood and lack known cellular readouts that can be used to assess the impact of genetic variation on protein function. In vitro, ubiquitin chain cleavage assays overcome this limitation as substrate-independent DUB activity readouts that can measure the impact of gene variants on enzymatic activity12.
In vitro ubiquitin cleavage assays have been used since the 1980s. These assays using radiolabeled substrates allowed for the discovery of the first DUBs, including isopeptidase, identified for its capacity to deubiquitylate Histone H2A13, and ubiquitin carboxyl-terminal hydrolase (UCH), identified by its ability to hydrolyze ubiquitin from a variety of chemical conjugates14,15,16. Further, radiolabeled polyubiquitylated full-length proteins or peptides were used to identify isopeptidase T and several UCHs and ubiquitin-specific proteases (UBPs) from erythrocytes and skeletal muscle, respectively17,18,19,20. Ubiquitin chains of a defined length and linkage type (K48-linked tetra-ubiquitin) were first used to measure the DUB activity of Isopeptidase T21. Since then, this assay has become the gold standard to measure DUB activity in mutational analyses22,23. The refinement of this assay currently allows visualization of ubiquitin cleavage via electrophoresis and conventional gel stains such as Coomassie blue, SYPRO orange, ruby, and silver stain or fluorescent or immunoblotting-based detection12,24. Molecular aspects of DUB activity, such as minimum chain length and linkage specificity25,26,27,28, can be clarified by using ubiquitin chains of different lengths (e.g., di-, tri-, tetra-ubiquitin) and linkages (K6, K11, K27, K29, K33, K48, K63, linear) in functional assays. NDD-associated variants can drive DUB activity defects that are ubiquitin chain linkage type specific11.
A di-ubiquitin cleavage assay using purified recombinant DUB proteins can directly measure the impact of NDD variants on DUB activity. USP27X, which is mutated in the NDD X-linked intellectual disability disorder 105 (XLID105)7,28 models the process presented here. This approach allows for the determination of how DUB activity is disrupted by gene variants in existing and unknown DUB-associated NDDs.
Protocol
The following protocol can be adapted for recombinant proteins using various affinity tags expressed in different strains of competent cells. Depending on the protein being expressed, the culturing and overnight expression conditions may require optimization of the OD600 at expression induction, expression time, expression temperature, and IPTG concentration. An overview of the protocol is illustrated in Figure 1. The details of the reagents and the equipment used in this study are listed in the Table of Materials.
1. Transformation of competent Rosetta 2 E. coli cells with the recombinant GST-USP27X expression plasmid
NOTE: To maintain the sterility of the bacterial culture, perform steps where media containers are open under a Bunsen burner flame. To allow optimal oxygen transfer, perform bacterial culture shaking in a benchtop temperature-controlled shaker with an orbit of 19 mm to 50 mm and a speed of 200 rpm29.
- Thaw 20 µL of chemically competent Rosetta 2 E. coli cells on ice immediately before use. Add 1-10 ng of the pGEX6P1-USP27X expression plasmid (encoding for N-terminal Glutathione-S-transferase (GST)30 -tagged USP27X and containing Ampicillin resistance) and incubate 5 min on ice.
- Heat shock for 30 s at 42 °C in a dry bath. Incubate on ice for 2 min.
- Add 80 µL of room temperature (RT) SOC medium. Incubate for 60 min at 37 °C with 200 rpm rotation in a 19 mm orbit benchtop temperature-controlled shaker.
- Plate 50 µL of culture on an LB agar plate supplemented with 25 µg/L Chloramphenicol and 50 µg/L Ampicillin. Incubate for 20 h at 37 °C lid-side down in a temperature-controlled incubator.
2. Overnight bacterial expression of recombinant protein from expression plasmid
NOTES: To maintain the sterility of the bacterial culture, perform steps where media containers are open under a Bunsen burner flame. To allow optimal oxygen transfer, perform bacterial culture shaking in a benchtop temperature-controlled shaker with an orbit of 19 mm to 50 mm and a speed of 200 rpm29. Measure culture OD600 using a spectrophotometer.
- Prepare 1 L of Terrific Broth (TB) medium by adding 47.6 g of TB Powder to 1 L of ultrapure water with 0.4% glycerol in a 2 L baffled culture flask. Autoclave medium for 30 min and cool to RT.
- Pick a single colony of transformed bacteria and add to 10 mL of sterile LB medium supplemented with 25 µg/L Chloramphenicol and 50 µg/L Ampicillin. Incubate for 20 h at 37 °C with 200 rpm rotation in a 19 mm orbit benchtop temperature-controlled shaker.
- Add 10 mL overnight culture to 1 L of TB medium supplemented with 25 µg/L Chloramphenicol and 50 µg/L Ampicillin (1:100 ratio of starting culture to expression culture). Incubate at 37 °C with 200 rpm rotation in a 19 mm orbit benchtop temperature-controlled shaker until the culture OD600 is between 0.5-0.6.
- Add 50 µM of IPTG to the culture to induce expression, cool to 16 °C, and incubate for 20 h at 16 °C with 200 rpm rotation.
- Pellet cells from expression culture by centrifugation for 20 min at ≥3,000 x g and 4 °C. Store the pellet at -80 °C for at least 1 h.
NOTE: At this point, the experiment can be paused and restarted later (preferably the same week). The pellet can be stored for the long term at -80 °C.
3. Protein purification by gravity-flow affinity column
NOTE: The resin, binding, wash, elution, and storage buffers appropriate for each purification will depend on the recombinant protein being purified. Collect samples from the cell pellet, supernatant, flow-through, wash fractions, and elution fractions in SDS-PAGE buffer. Perform SDS-PAGE and Coomassie staining for the samples to evaluate the success of the purification. Perform purification at 4 °C and handle fractions on ice. Cleavage of protein tags can be performed on or off the column using the appropriate protease to target the relevant protease-specific cleavage site.
- Secure the empty gravity flow column in the retort stand and fill it with glutathione agarose resin. Use 2-3 mL of resin to purify a cell pellet collected from 1 L of expression culture.
- Wash the column with one resin-bed volume of 20% ethanol. Wash column 3 times with one resin-bed volume of MS500 buffer (20 mM of Tris pH 7.5, 500 mM of NaCl, 0.5 mM of TCEP). Stop the column so the resin remains covered with buffer to prevent drying while preparing the cell pellet.
- Thaw the cell pellet at 4 °C. Add 30 mL of MS500 buffer (20 mM of Tris pH 7.5, 500 mM of NaCl, 0.5 mM of TCEP, 60 mg of lysozyme, and one protease inhibitor tablet) to the thawed pellet. Lyse for 30 min at 4 °C with gentle end-over-end rotation.
- Sonicate lysed cells in either a 50 mL centrifuge tube or a metal beaker on ice until the lysate flows freely when dispensed from a pipette tip. Centrifuge for 30 min at 4 °C and ≥20,000 x g to clear the supernatant.
NOTE: Determine sonicator settings empirically. Set the sonicator such that 120 s total time is enough to reduce the viscous lysate to a free-flowing and transparent liquid. - Decant the supernatant into a beaker and load cleared lysate onto the column. Run lysate through the column by gravity flow while collecting the flow through. Load the column with the collected flow through and run it through the column.
- Wash the column with at least two resin-bed volumes of MS500 wash buffer (20 mM of Tris pH 7.5, 500 mM of NaCl, 0.5 mM of TCEP) 5 times. Collect the wash flow through in 5 mL fractions. Add 1 µL of each fraction to 100 µL of Bradford reagent to visually check for protein presence. Wash until unbound protein is no longer present in the last wash, adding additional wash steps if necessary.
- Recover protein by running MS500 elution buffer (MS500 buffer supplemented with 10 mM of glutathione and 10 mM of NaOH) through the column, collecting 5 mL elution fractions. Check for protein presence by adding 1 µL of eluate to 100 µL of Bradford reagent. Stop collecting fractions when the Bradford reagent no longer indicates proteins are present.
- Perform buffer exchange by precipitation and centrifugation. Precipitate protein by adding 2 volumes of 4 M of ammonium sulfate to the eluate, gently inverting until cloudy, and then centrifuging for 30 min at 4 °C and ≥20,000 x g and again for another 5 min. Remove the supernatant after each centrifugation without disturbing the protein pellet.
- Re-dissolve and store protein in MS500 supplemented with 25% glycerol. Store protein pellets or protein in storage buffer at -80 °C.
4. In vitro ubiquitin chain cleavage assay
NOTE: Select ubiquitin chain length and linkage types based on DUB specificity described in previous reports31 or determined empirically. If necessary, this protocol could be used to test the activity of the wild-type DUB of interest on a panel of commercially available ubiquitin chains of defined length and linkage type. A di-ubiquitin chain amount of 375-750 ng and a DUB concentration of 1-2 µM can be used as starting points for the assay27.
- Prepare 10x DUB activation buffer (500 mM of Tris-HCl pH 7.5, 500 mM of NaCl, and 100 mM of TCEP).
- For each time point for each DUB, prepare 10 µL total of 2 µM of purified GST-USP27X in 1x DUB activation buffer (DUB mix). Prepare master mixes and split them into time points.
- Incubate the DUB mix for 10 min at RT.
- Add 7 µL of SDS-PAGE loading buffer to time 0 before adding ubiquitin chains to prevent the deubiquitylation reaction from starting.
- To each time point for each DUB, add 375 ng of K-63 di-ubiquitin chains diluted in 10 µL 1x DUB activation buffer. The total volume is 20 µL per reaction.
- Incubate the tubes at 30 °C, stopping each time point with 7 µL of SDS-PAGE loading buffer.
- Perform SDS-PAGE with a 4%-12% gradient gel and immunoblot7,32 for ubiquitin and USP27X to analyze the change in mono-ubiquitin presence across selected time points.
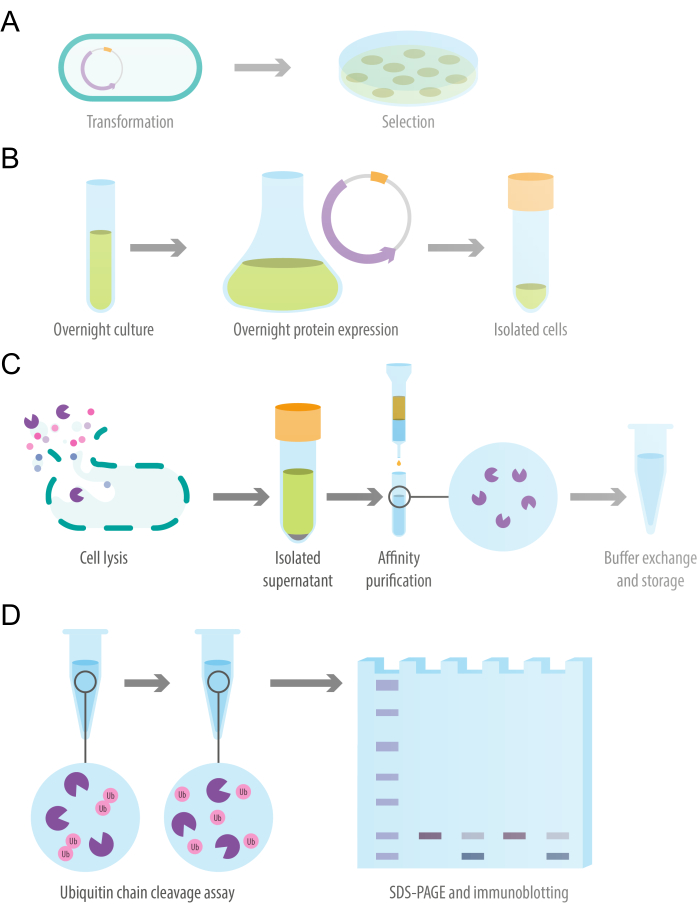
Figure 1: Schematic of the study design. (A) Transformation of competent E. coli cells with recombinant protein expression plasmid. (B) Overnight bacterial expression of recombinant deubiquitylase protein. (C) Protein purification of recombinant deubiquitylase using a gravity-flow affinity column. (D) In vitro ubiquitin chain cleavage assay to evaluate deubiquitylating activity. Please click here to view a larger version of this figure.
Results
To determine the impact of XLID105-associated variants on USP27X catalytic activity, GST-tagged wild-type USP27X and the XLID105-associated variant F313V, Y381H, and S404N USP27X proteins were purified from bacteria. These variants are located within the USP catalytic domain of USP27X (Figure 2A). Because USP27X was previously reported to cleave K63 ubiquitin chains31, wild-type USP27X and the XLID105-associated variant F313V, Y381H, and S404N USP27X proteins were inc...
Discussion
This article presents a protocol for the expression and purification of recombinant USP27X DUBs and an in vitro ubiquitin chain cleavage assay to compare the deubiquitylating activity of wild-type USP27X and NDD-associated variant proteins. This assay determined that XLID105-associated variants disrupt USP27X catalytic activity7. This mechanistic insight helped us define XLID105 as a USP27X functional disruption disorder.
This protocol can be adapted to other D...
Disclosures
The authors declare no conflicts of interest.
Acknowledgements
This work was funded by Sanford Research startup funds to FB and the NIH grant R01CA233700 to MJS. The artwork was done by Felipe G. Serrano (www.illustrative-science.com). We thank Dr. Greg Findlay (University of Dundee) for the GST-USP27X plasmid.
Materials
| Name | Company | Catalog Number | Comments |
| Amersham Protran 0.45 NC 200 mm × 200 mm 25/PK | Cytiva | 10600041 | |
| Ammonium sulfate | Fisher Scientific | AC205872500 | |
| Ampicillin | Fisher Scientific | BP1760 25 | |
| Anti- Ubiquitin (Mouse monoclonal) | Biolegend | Cat# 646302, RRID:AB_1659269 | (WB: 1:1000) |
| Anti-GST (Sheep polyclonal) | MRC-PPU Reagents and Services | Cat# S902A Third bleed | (WB: 1:1000) https://mrcppureagents.dundee.ac.uk/ |
| Baffled Culture Flasks 2 L | Fisher Scientific | 10-042-5N | |
| Bradford Reagent | Millipore Sigma | B6916-500ML | |
| Chloramphenicol | Gold Biotechnology | C-105-25 | |
| Complete, Protease Inhibitor tablets | Millipore Sigma | 5056489001 | |
| Econo-Column 1.5 × 5 cm | Bio-Rad | 7371507 | |
| Eppendorf ThermoMixer F1.5 | Eppendorf | 5384000020 | |
| Excel | Microsoft | https://www.microsoft.com/en-us/microsoft-365/excel | |
| Glycerol | Genesee Scientific | 18-205 | |
| Illustrator | Adobe | https://www.adobe.com/products/illustrator.html | |
| Image Studio | LI-COR Biosciences | https://www.licor.com/bio/image-studio/ | |
| Inkscape | Inkscape | https://inkscape.org/ | |
| Invitrogen 4-12% NuPAGE 1mm 12 well gel | Thermo Fisher Scientific | NP0322BOX | |
| IPTG (Isopropyl-b-D-Thiogalactopyranoside) | Genesee Scientific | 20-109 | |
| IRDye 800CW Donkey anti-Goat IgG Secondary Antibody | LI-COR Biosciences | Cat# 926-32214 | (WB: 1:10000) |
| IRDye 800CW Donkey anti-Mouse IgG Secondary Antibody | LI-COR Biosciences | Cat# 926-32212 | (WB: 1:10000) |
| Isotemp Digital Dry Bath | Fisher Scientific | 88860022 | |
| K63 Di-Ubiquitin | South Bay Bio LLC | SBB-UP0072 | |
| LB Agar | Genesee Scientific | 11-119 | |
| LB Broth | Genesee Scientific | 11-118 | |
| Lysozyme | Gold Biotechnology | L-040-100 | |
| MaxQ 4000 Benchtop Orbital Shaker | Thermo Fisher Scientific | SHKE4000-7 | |
| MES-SDS Running Buffer | Boston Bioproducts Inc | BP-177 | |
| Mini Tube Rotator | Fisher Scientific | 88-861-051 | |
| NuPage LDS Sample buffer 4x | Thermo Fisher Scientific | NP0007 | |
| Odyssey Fc Imager | LI-COR Biosciences | 43214 | |
| PageRuler Plus Ladder | Thermo Fisher Scientific | 26620 | |
| pGEX6P1 human USP27X | MRC-PPU Reagents and Services | DU21193 | https://mrcppureagents.dundee.ac.uk/ |
| pGEX6P1 human USP27X F313V | Addgene | 225715 | Koch et at 2024 (PMID: 38182161) |
| pGEX6P1 human USP27X S404N | Addgene | 225717 | Koch et at 2024 (PMID: 38182161) |
| pGEX6P1 human USP27X Y381H | Addgene | 225716 | Koch et at 2024 (PMID: 38182161) |
| Pierce Glutathione Agarose | Thermo Fisher Scientific | 16100 | |
| PMSF (Phenylmethylsulfonyl fluoride) | Gold Biotechnology | P-470-10 | |
| Polysorbate 20 (Tween 20) | Fisher Scientific | AC233360010 | |
| Rosetta 2 Competent Cells | Millipore Sigma | 71402-M | |
| SimplyBlue SafeStain | Thermo Fisher Scientific | LC6060 | |
| SmartSpec 3000 | Bio-Rad | 170-2501 | |
| SOC medium | Thermo Fisher Scientific | 15544034 | |
| Sodium chloride | Genesee Scientific | 18-216 | |
| Sonifier 250 | Branson | 100-132-135 | |
| Sorvall RC 6 Plus Centrifuge | Thermo Fisher Scientific | 46910 | |
| TCEP (Tris-(carboxyethyl) phosphine hydrochloride) | Gold Biotechnology | TCEP10 | |
| Terrific Broth Powder | Genesee Scientific | 18-225 | |
| Tris Base | Genesee Scientific | 18-146 | |
| XCell SureLock Mini-Cell and XCell II Blot Module | Thermo Fisher Scientific | EI0002 |
References
- Morris-Rosendahl, D. J., Crocq, M. -. A. Neurodevelopmental disorders: the history and future of a diagnostic concept. Dialogues Clin Neurosci. 22 (1), 65-72 (2020).
- Ebstein, F., Küry, S., Papendorf, J. J., Krüger, E. Neurodevelopmental disorders (NDD) caused by genomic alterations of the ubiquitin-proteasome system (UPS): the possible contribution of immune dysregulation to disease pathogenesis. Front Mol Neurosci. 14, 733012 (2021).
- Hershko, A., Ciechanover, A. The ubiquitin system. Annu Rev Biochem. 67 (10), 425-479 (1998).
- Pickart, C. M. Mechanisms underlying ubiquitination. Annu Rev Biochem. 70 (1), 503-533 (2001).
- Komander, D., Clague, M. J., Urbé, S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 10 (8), 550-563 (2009).
- Yau, R., Rape, M. The increasing complexity of the ubiquitin code. Nat Cell Biol. 18 (6), 579-586 (2016).
- Koch, I., et al. USP27X variants underlying X-linked intellectual disability disrupt protein function via distinct mechanisms. Life Sci Alliance. 7 (3), e202302258 (2024).
- Hao, Y. -. H., et al. USP7 acts as a molecular rheostat to promote WASH-dependent endosomal protein recycling and is mutated in a human neurodevelopmental disorder. Mol Cell. 59 (6), 956-969 (2015).
- Fountain, M. D., et al. Pathogenic variants in USP7 cause a neurodevelopmental disorder with speech delays, altered behavior, and neurologic anomalies. Genet Med. 21 (8), 1797-1807 (2019).
- Santiago-Sim, T., et al. Biallelic variants in OTUD6B cause an intellectual disability syndrome associated with seizures and dysmorphic features. Am J Hum Genet. 100 (4), 676-688 (2017).
- Beck, D. B., et al. Linkage-specific deubiquitylation by OTUD5 defines an embryonic pathway intolerant to genomic variation. Sci Adv. 7 (4), eabe2116 (2021).
- Cho, J., Park, J., Kim, E. E., Song, E. J. Assay systems for profiling deubiquitinating activity. Int J Mol Sci. 21 (16), 1-16 (2020).
- Matsui, S., Sandberg, A. A., Negoro, S., Seon, B. K., Goldstein, G. Isopeptidase: A novel eukaryotic enzyme that cleaves isopeptide bonds. Proc Natl Acad Sci USA. 79 (5), 1535-1539 (1982).
- Rose, I. A., Warms, J. V. B. An enzyme with ubiquitin carboxy-terminal esterase activity from reticulocytes. Biochemistry. 22 (18), 4234-4237 (1983).
- Wilkinson, K. D., Cox, M. J., Mayer, A. N., Frey, T. Synthesis and characterization of ubiquitin ethyl ester, a new substrate for ubiquitin carboxyl-terminal hydrolase. Biochemistry. 25 (21), 6644-6649 (1986).
- Pickart, C. M., Rose, I. A. Ubiquitin carboxyl-terminal hydrolase acts on ubiquitin carboxyl-terminal amides. J Biol Chem. 260 (13), 7903-7910 (1985).
- Hadari, T., Warms, J. V. B., Rose, I. A., Hershko, A. A ubiquitin C-terminal isopeptidase that acts on polyubiquitin chains: Role in protein degradation. J Biol Chem. 267 (2), 719-727 (1992).
- Woo, S. K., et al. Multiple ubiquitin C-terminal hydrolases from chick skeletal muscle. J Biol Chem. 270 (32), 18766-18773 (1995).
- Woo, S. K., et al. Purification and characterization of a new ubiquitin C-terminal hydrolase (UCH-1) with isopeptidase activity from chick skeletal muscle. J Biochem. 121 (4), 684-689 (1997).
- Baek, S. H., et al. Molecular cloning of a novel ubiquitin-specific protease, UBP41, with isopeptidase activity in chick skeletal muscle. J Biol Chem. 272 (41), 25560-25565 (1997).
- Wilkinson, K. D., et al. Metabolism of the polyubiquitin degradation signal: Structure, mechanism, and role of isopeptidase T. Biochemistry. 34 (44), 14535-14546 (1995).
- Hu, M., et al. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell. 111 (7), 1041-1054 (2002).
- Hu, M., et al. Structure and mechanisms of the proteasome-associated deubiquitinating enzyme USP14. EMBO J. 24 (21), 3747-3756 (2005).
- Gorka, M., Magnussen, H. M., Kulathu, Y. Chemical biology tools to study deubiquitinases and UBL proteases. Semin Cell Dev Biol. 132, 86-96 (2022).
- Faesen, A. C., et al. The differential modulation of USP activity by internal regulatory domains, interactors, and eight ubiquitin chain types. Chem Biol. 18 (12), 1550-1561 (2011).
- Mevissen, T. E. T., et al. OTU deubiquitinases reveal mechanisms of linkage specificity and enable ubiquitin chain restriction analysis. Cell. 154 (1), 169-184 (2013).
- Kwasna, D., et al. Discovery and characterization of ZUFSP/ZUP1, a distinct deubiquitinase class important for genome stability. Mol Cell. 70 (1), 150-164.e6 (2018).
- Hu, H., et al. X-exome sequencing of 405 unresolved families identifies seven novel intellectual disability genes. Mol Psychiatry. 21 (1), 133-148 (2016).
- Shaker Orbit - Revolving in Space Around the Samples? Eppendorf Lab Academy. Eppendorf Lab Academy Available from: https://www.eppendorf.com/us-en/lab-academy/lab-solutions/others/shaker-orbit-revolving-in-space-around-the-samples (2020)
- Smith, D. B., Johnson, K. S. Single-step purification of polypeptides expressed in Escherichia coli as fusions with glutathione S-transferase. Gene. 67 (1), 31-40 (1988).
- Ritorto, M. S., et al. Screening of DUB activity and specificity by MALDI-TOF mass spectrometry. Nat Commun. 5, 4763 (2014).
- Wingfield, P. T. Overview of the purification of recombinant proteins. Curr Protoc Protein Sci. 80, 6.1.1-6.1.35 (2015).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved